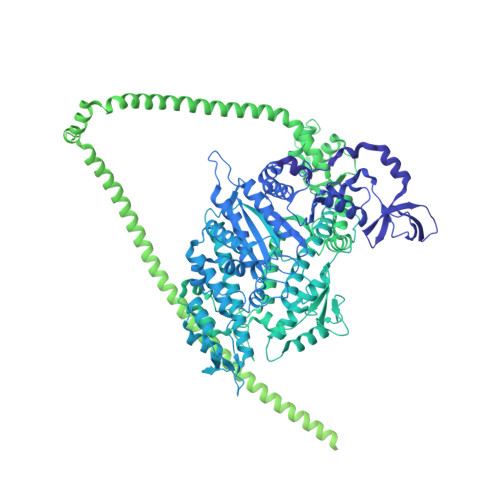
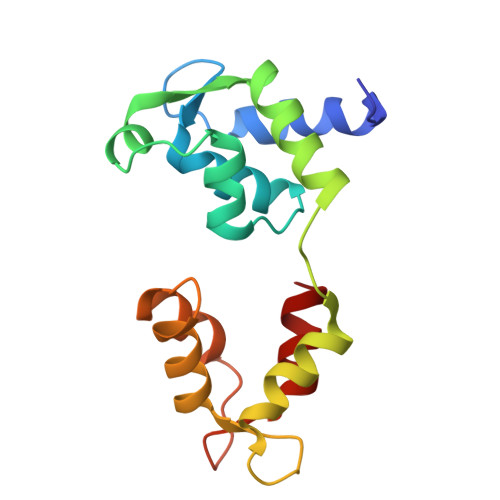
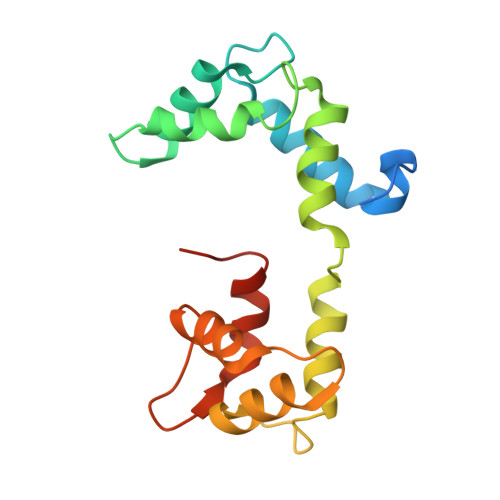
Cryo-EM structure of shutdown human nonmuscle myosin 2A.
Casas-Mao, D., Carrington, G., Peckham, M.(2026) Sci Adv 12: eaed1858-eaed1858
- PubMed: 41996489 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.aed1858
- Primary Citation Related Structures:
9SYU, 9SZR - PubMed Abstract:
Determining the high-resolution structure of the widely expressed nonmuscle myosin 2A (NM2A), in its dephosphorylated shutdown state, is important in understanding its regulation and disease roles. In shutdown molecules, the coiled-coil tail wraps around the myosin heads, preventing them from forming filaments and binding to actin. We have solved the shutdown structure of NM2A to a global resolution of 3.0 angstroms in the head region and 6.3 angstroms for the whole molecule. This reveals specific ionic interactions that explain why the path of the coiled coil and the shutdown mechanism for NM2A differ from those of β-cardiac myosin and provides key insight into how specific mutations likely destabilize the shutdown state, leading to disease.
- Astbury Centre for Structural Molecular Biology & School of Molecular and Cellular Biology, Faculty of Biological Sciences, University of Leeds, Leeds, UK.
Organizational Affiliation: