Discovery of Potent, Selective, and Brain-Penetrant Small Molecule CD38 Inhibitors.
Stott, A.J., Burli, R.W., Doyle, K.J., Dickson, L., Hewer, R.C., Pickford, P., Roberts, M.J., Waters-Hall, R., Wu, Y., Zebisch, M., Rangel, V., Geitmann, M., Matthews, K., Brice, N.L., Carlton, M., Dawson, L.A., Harvey, J.R.M.(2026) ACS Med Chem Lett 17: 259-265
- PubMed: 41704394
- DOI: https://doi.org/10.1021/acsmedchemlett.5c00727
- Primary Citation Related Structures:
9SXI - PubMed Abstract:
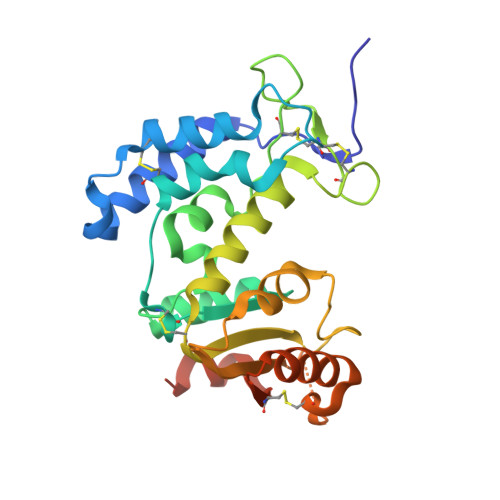
Cluster of differentiation 38 (CD38) is a nicotinamide adenine dinucleotide (NAD + )-consuming ectoenzyme abundantly expressed in brain regions involved in motor control and cognition. Given the central role of NAD + in maintaining neuronal health, inhibition of CD38, resulting in NAD + elevation, has emerged as a potential therapeutic approach for neurodegenerative diseases and age-associated cognitive decline. Herein, we report the rational, structure-guided optimization of a series of small-molecule CD38 inhibitors, culminating in the identification of CVN14 , a potent, selective, and brain-penetrant tool molecule with favorable pharmacokinetic properties for advanced preclinical evaluation. We further disclose the first high-resolution X-ray crystal structure of the CVN14- ADPR - CD38 complex, revealing an uncompetitive binding mode. CVN14 provides a molecular tool to investigate CD38 biology in neurodegeneration and supports the development of next-generation brain-penetrant CD38 inhibitors.
- Cerevance Limited, 418 Cambridge Science Park, Cambridge CB4 0PZ, U.K.
Organizational Affiliation: