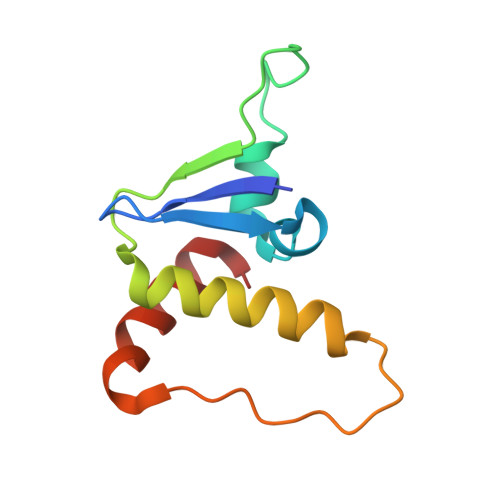
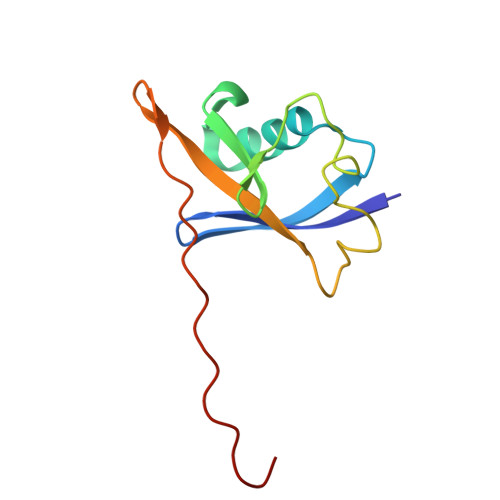
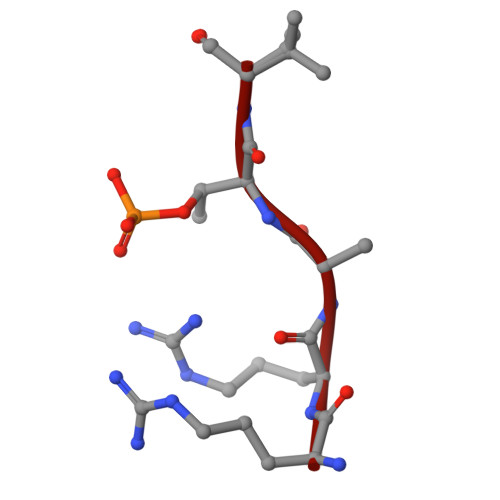
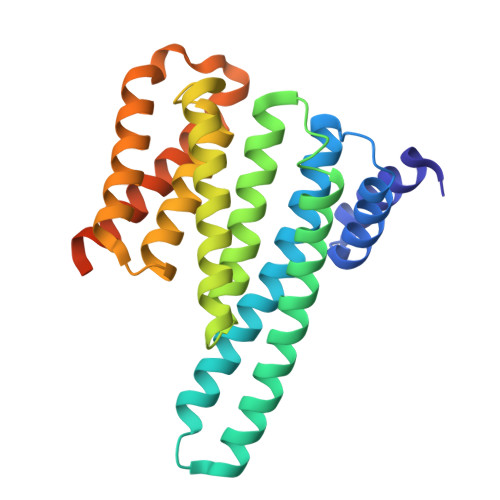
Induced ubiquitination of the partially disordered Estrogen Receptor alpha protein via a 14-3-3-directed molecular glue-based PROTAC design
Verhoef, C.J.A., Crowe, C., Nakasone, M.A., DeLaCuadra-Baste, A., Harzing, T., Span, N.A.S., Sathe, G., Iso, K., Ottmann, C., Brunsveld, L., Ciulli, A., Cossar, P.J.(2025) bioRxiv