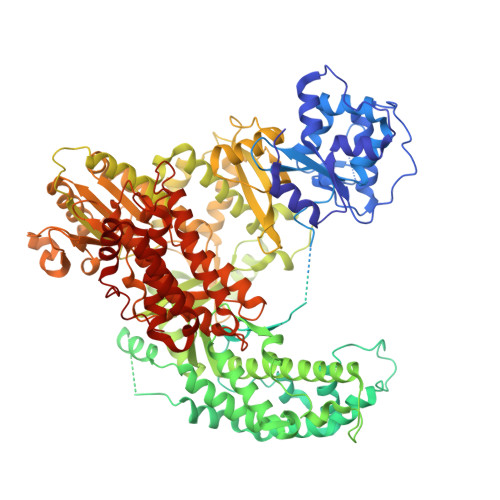
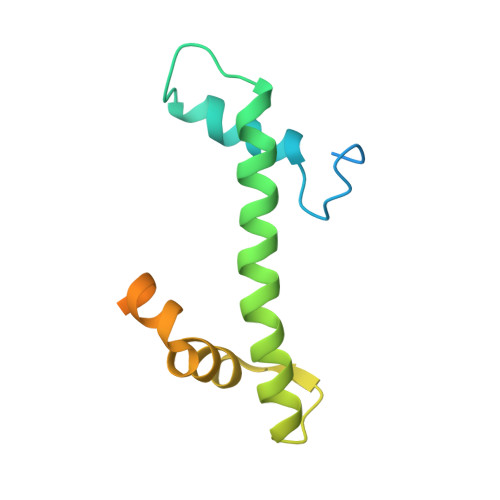
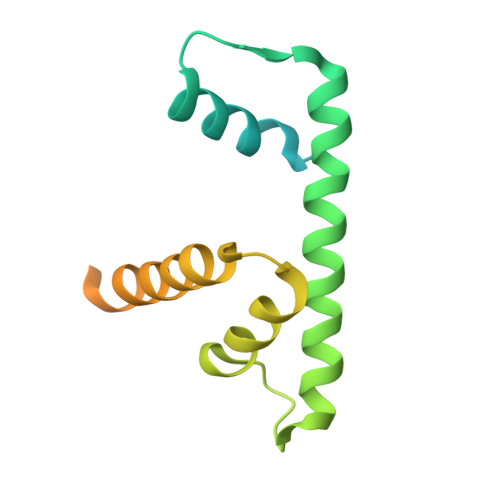
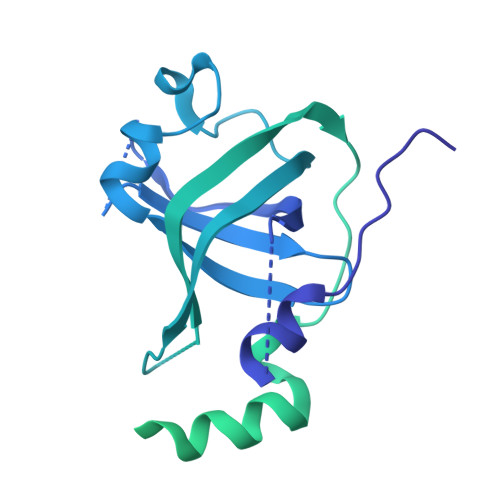
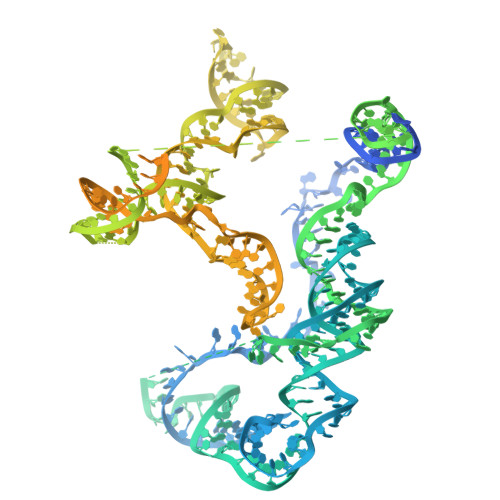
Structures of nucleotide-bound human telomerase at several steps of its telomeric DNA repeat addition cycle.
Balch, S., Franco-Echevarria, E., Ghanim, G.E., Kretsch, R.C., Das, R., Nguyen, T.H.D.(2026) Nat Commun 17: 1847-1847
- PubMed: 41565648 Search on PubMed
- DOI: https://doi.org/10.1038/s41467-026-68560-8
- Primary Citation Related Structures:
9SHY, 9SHZ, 9SI0 - PubMed Abstract:
In most eukaryotes, the reverse transcriptase telomerase counteracts telomere shortening by processively adding telomeric DNA repeat sequences to chromosome ends. Telomerase activity depends on the telomerase reverse transcriptase (TERT) and the telomerase RNA (hTR in humans). Processive telomere elongation is critical for genome stability, and defects in this mechanism are linked to cellular dysfunction and human disease. However, the structural basis for telomerase repeat addition processivity in humans has remained elusive. Here, we present cryo-electron microscopy structures of human telomerase bound to telomeric DNA and an incoming nucleotide, captured at three distinct stages of its repeat addition cycle: initiation, elongation, and pre-termination. Across these states, the TERT active site maintains a conserved architecture that stabilises a short DNA-RNA duplex of constant length of four base-pairs. Beyond the active site, we identify dynamic structural features in both TERT and hTR that facilitate substrate engagement and RNA template repositioning, thereby supporting the synthesis of successive telomeric repeats. Together, these structures provide key insights into how human telomerase achieves its unique processivity to maintain telomere length and genome integrity.
- Medical Research Council Laboratory of Molecular Biology, Francis Crick Avenue, Cambridge, UK.
Organizational Affiliation: