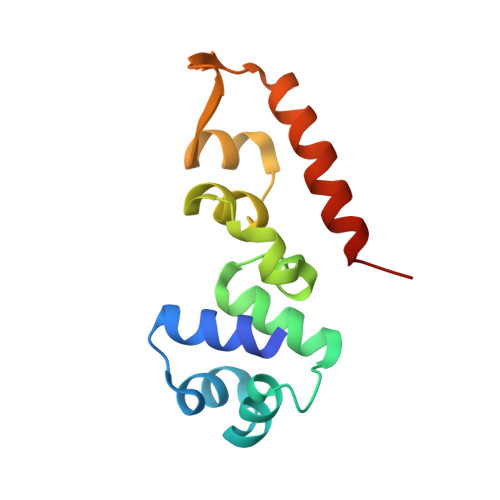
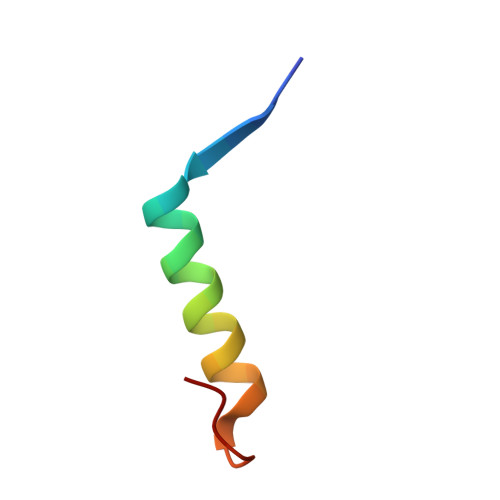
A widespread extended arbitrium system controls lysis/lysogeny through antirepression.
Kabel, S., Omer Bendori, S., Borenstein, T., Guler, P., Martinez-Alonso, C., Mancheno-Bonillo, J., Gallego-Del-Sol, F., Marina, A., Eldar, A.(2026) Cell Host Microbe 34: 278
- PubMed: 41619737 Search on PubMed
- DOI: https://doi.org/10.1016/j.chom.2026.01.007
- Primary Citation Related Structures:
9IAQ, 9SF2 - PubMed Abstract:
Many temperate Bacillus phages use the arbitrium peptide-based signaling system to regulate lysis-lysogeny decisions. In this system, the secreted AimP peptide inhibits the AimR receptor to promote lysogeny. However, the downstream mechanism of AimR-mediated lysis control remains unclear for most systems. Here, we identify that ∼75% of arbitrium systems possess an extended five-gene module, including the aimX, aimC, and aimL genes. AimX encodes a small AimR-regulated antirepressor protein that binds the phage repressor AimC, preventing its oligomerization and DNA binding, thereby activating the pro-lytic aimL gene and additional lytic genes. This mechanism was validated across multiple phages and structurally characterized, revealing that AimX mimics the AimC oligomerization domain to prevent oligomerization and inhibit repressor function. These findings elucidate the predominant molecular strategy by which arbitrium systems control phage lysis-lysogeny transitions and highlight the central role of small proteins in phage decision-making.
- Shmunis School of Biomedicine and Cancer Research, Faculty of Life Sciences, Tel Aviv University, 6139001 Tel Aviv, Israel.
Organizational Affiliation: