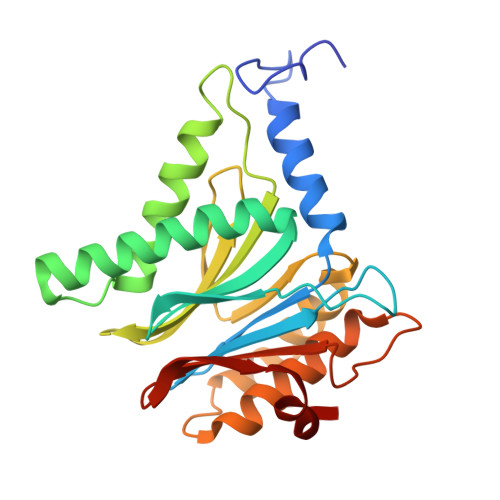
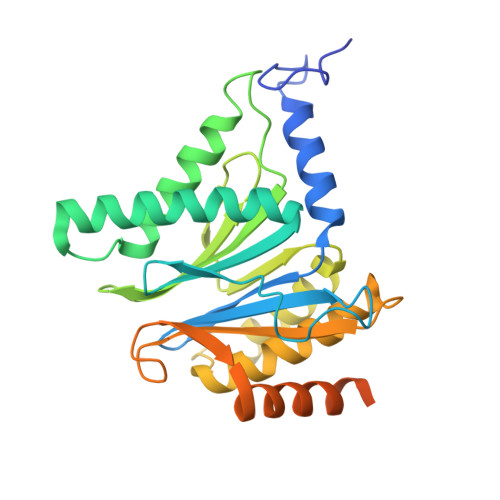
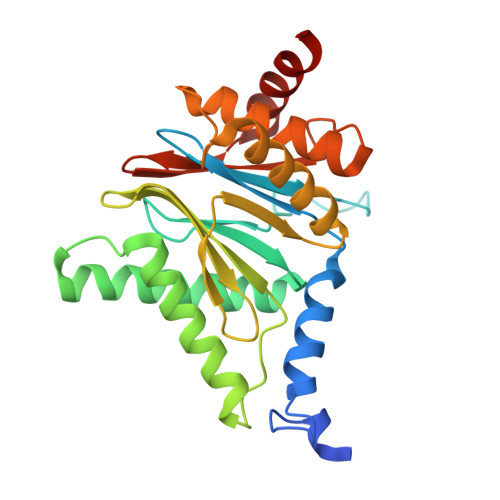
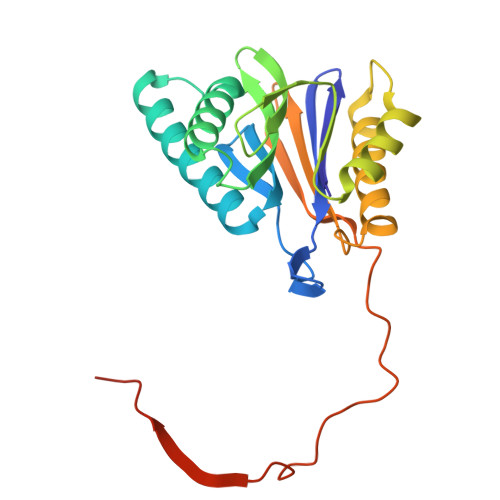
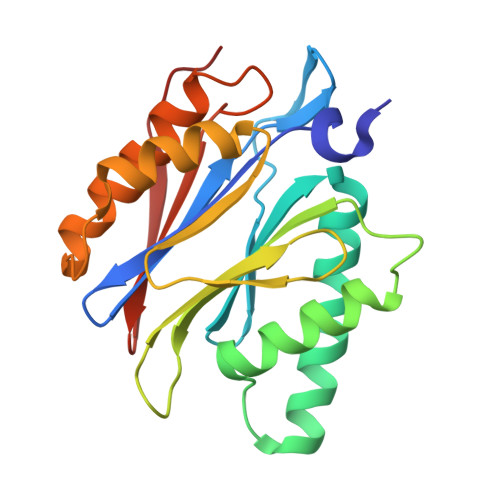
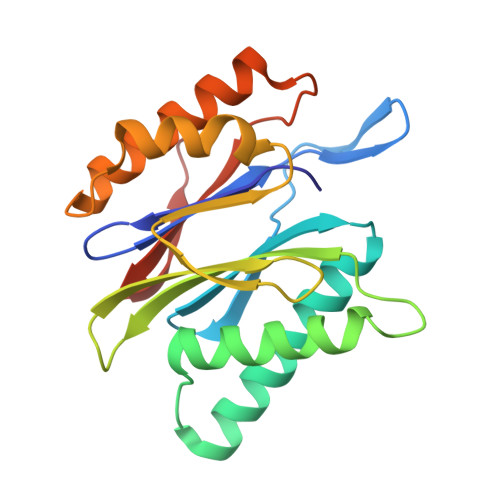
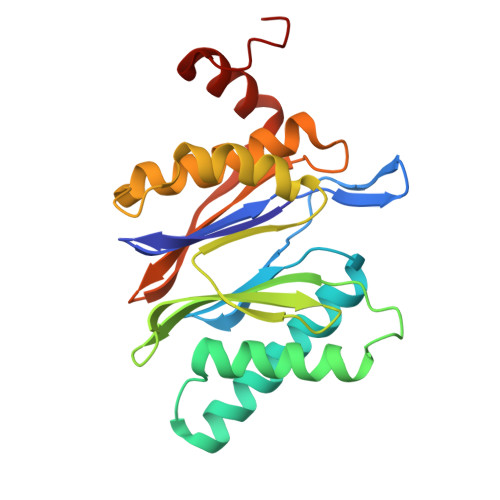
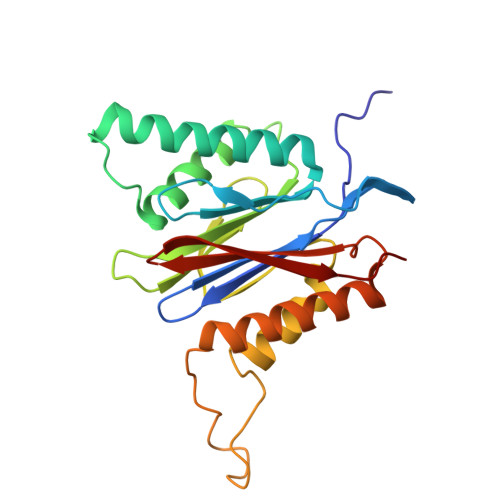
Tellurophene-Tagged Carfilzomib Enables Single-Cell Mass Cytometric Mapping of Proteasome Activity.
Potter, N., Eddenden, A., Fomina, A., Dinesh, A., Jackson, H.W., McGuigan, A.P., Groll, M., Nitz, M.(2025) ACS Chem Biol 20: 2936-2942
- PubMed: 41252648
- DOI: https://doi.org/10.1021/acschembio.5c00691
- Primary Citation of Related Structures:
9S78 - PubMed Abstract:
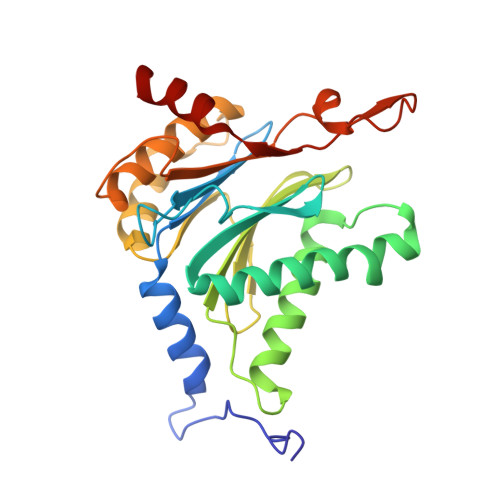
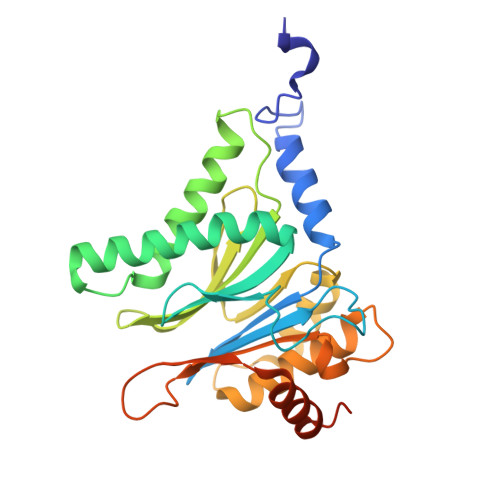
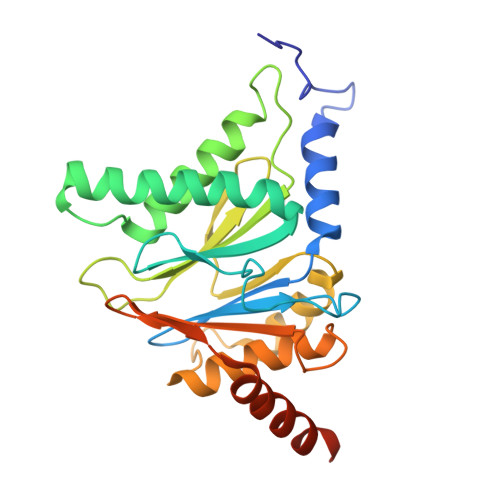
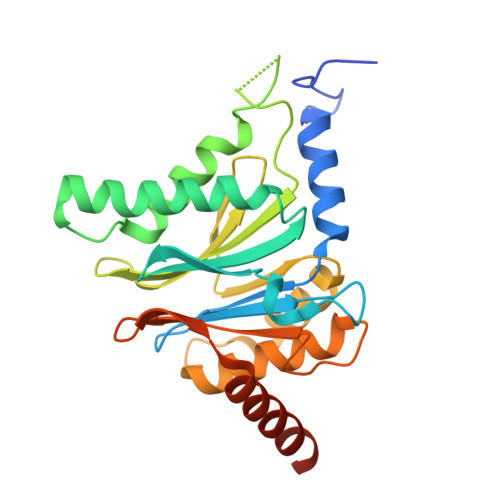
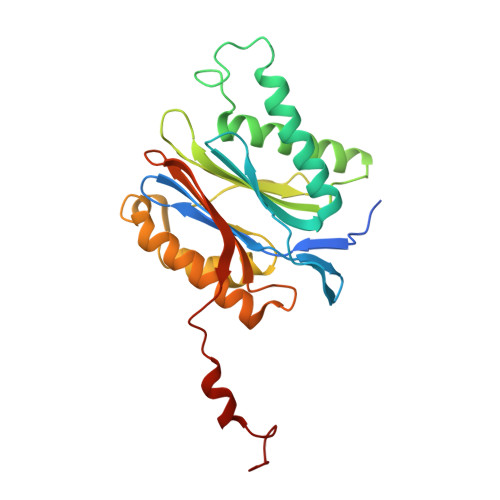
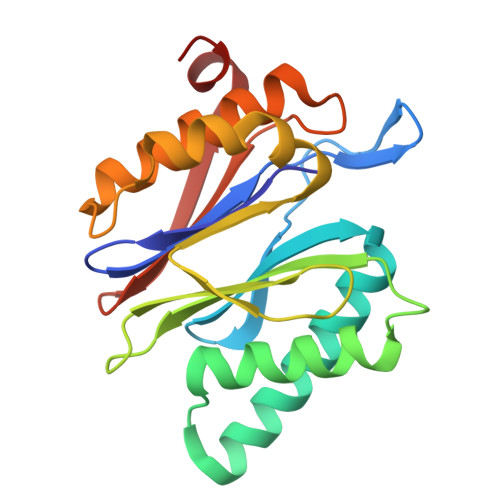
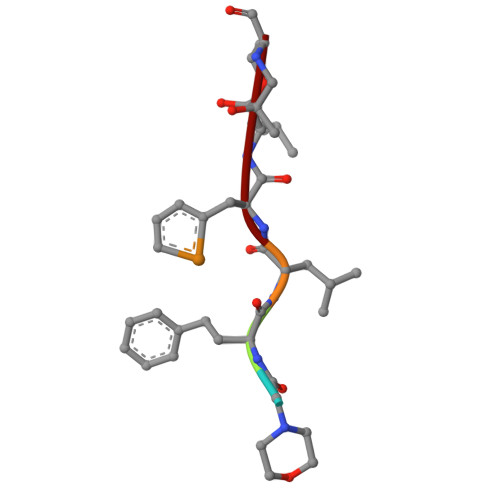
Tracking small-molecule distribution in heterogeneous cell samples at single-cell resolution remains a major analytical challenge. Here, we present a tellurophene-functionalized analogue of the proteasome inhibitor Carfilzomib (TeCar) whose distribution can be followed by mass cytometric (MC) quantification while preserving target engagement and cytotoxicity. Structural and biochemical analyses confirm that TeCar binds the proteasome in a mode comparable to the clinically approved parent compound. Using MC, we demonstrate selective TeCar accumulation in malignant over immune cells within mixed populations, with cancer cells exhibiting 15 to 30-fold higher uptake. Tellurium signal correlates with proteasomal activity, and differential labeling among immune subsets reveals functional heterogeneity not captured by transcriptomics alone. These findings establish tellurophene tagging as a minimally perturbing and broadly applicable strategy for functional distribution studies at single-cell resolution.
- Department of Chemistry, University of Toronto, Toronto M5S3H6, Canada.
Organizational Affiliation: