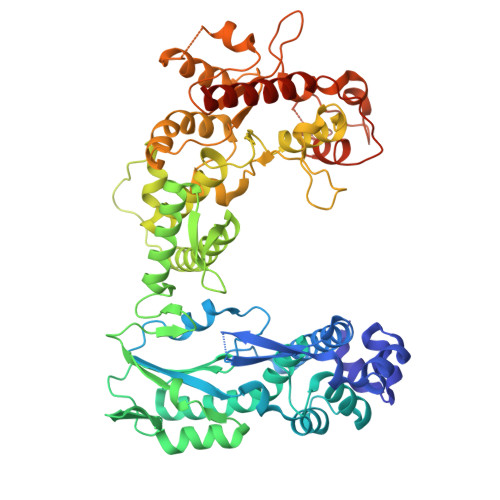
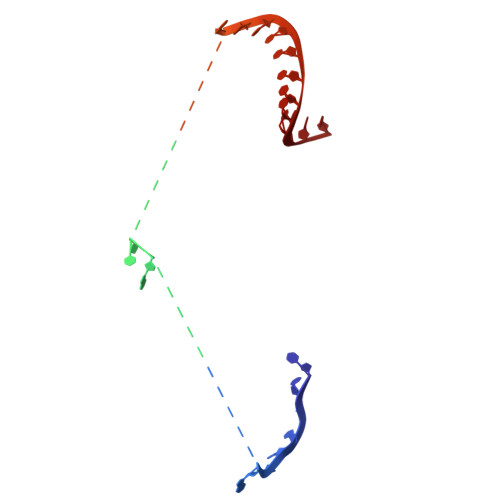
Structure and mechanism of antiphage retron Eco2.
Jasnauskaite, M., Juozapaitis, J., Liegute, T., Grigaitis, R., Skorupskaite, A., Steinchen, W., Miksys, A., Truncaite, L., Kazlauskaite, K., Torres Jimenez, M.F., Khochare, S., Dudas, G., Bange, G., Malinauskaite, L., Songailiene, I., Pausch, P.(2026) Nat Struct Mol Biol 33: 330-340
- PubMed: 41709047 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-026-01754-2
- Primary Citation Related Structures:
9I2F, 9I2G, 9S1F - PubMed Abstract:
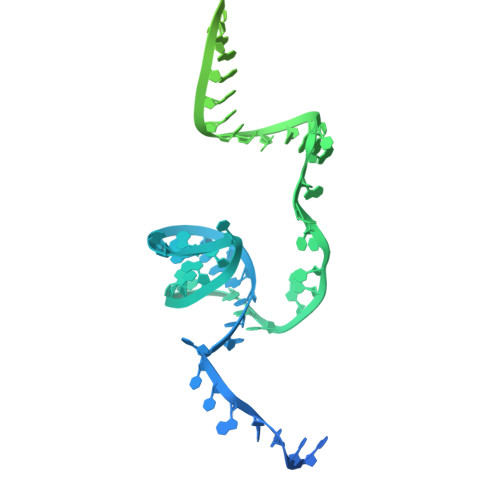
Retrons are prokaryotic reverse transcriptase systems that produce multicopy single-stranded DNA (msDNA), yet the principles by which they mediate antiviral defense remain largely unresolved. Here we investigate the mechanism of Escherichia coli Eco2, a minimal retron composed of a single reverse transcriptase-nuclease fusion protein. Cryogenic electron microscopy and hydrogen/deuterium exchange mass spectrometry reveal the structures and dynamics of a trimeric nucleoprotein complex assembled within a branched msDNA scaffold, which cages the TOPRIM nucleases. We show that the phage-encoded endonuclease DenB initiates msDNA degradation, thereby unblocking the nuclease active sites. Activated Eco2 cuts transfer RNAs, resulting in translational shutdown for antiphage defense. We further identify ribosomal protein S1 as a putative RNA chaperone that associates with the msDNA precursor. These findings provide insights into the molecular mechanisms of minimal retrons and establish a structural basis for engineering of Eco2.
- LSC-EMBL Partnership Institute for Genome Editing Technologies, Life Sciences Center, Vilnius University, Vilnius, Lithuania.
Organizational Affiliation: