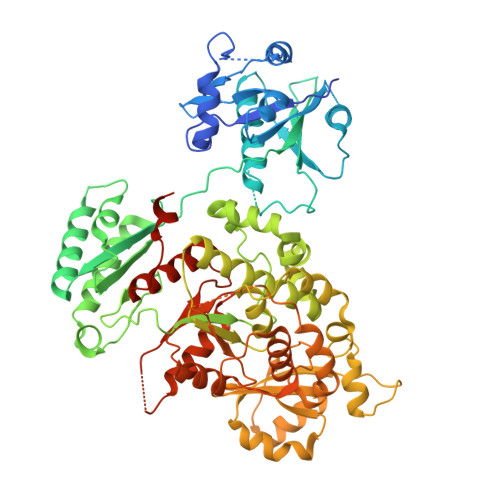
Prokaryotic PfaB is a terminal acyltransferase that determines the final PUFA product.
Lofeudo, N., Martin, A., Jacome, M., Wan, X., Lucas, M., Moncalian, G.(2026) Protein Sci 35: e70497-e70497
- PubMed: 41676921 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.70497
- Primary Citation Related Structures:
9RY8 - PubMed Abstract:
Omega-3 polyunsaturated fatty acids (PUFAs) are essential for human health due to their numerous beneficial biological properties. These compounds are synthesized in marine bacteria and eukaryotic microalgae by PUFA megasynthases (Pfas), which are evolutionarily related to fatty acid synthases (FAS) and polyketide synthases (PKS). In FAS, PKS, and PUFA synthases, the acyltransferase (AT) domain plays a critical role in condensation reactions by loading starter or extender units into the acyl carrier protein (ACP) domain. PfaB, a component of PUFA megasynthases, harbors a pseudo-ketosynthase (KS') domain and an AT domain. In this study, we show that PfaB determines the final PUFA product, as demonstrated by in vivo assays in Escherichia coli using the DHA-producing Moritella marina and the EPA-producing Shewanella baltica. In vitro biochemical assays confirm that PfaB exhibits acyltransferase activity, with distinct substrate specificity from the AT domain of PfaA. Finally, we report the crystal structure of PfaB from S. baltica, representing the first structurally resolved AT domain within a PUFA megasynthase. Molecular docking analyses suggest that specific residues may contribute to differences in substrate recognition and specificity. Together, these findings show that PfaB acts as the terminal acyltransferase, providing new insights into its functional role in PUFA biosynthesis, and advancing our understanding of its mechanism and ligand interactions.
- Department of Molecular Biology, Institute of Biomedicine and Biotechnology of Cantabria (IBBTEC), University of Cantabria-CSIC, Santander, Spain.
Organizational Affiliation: