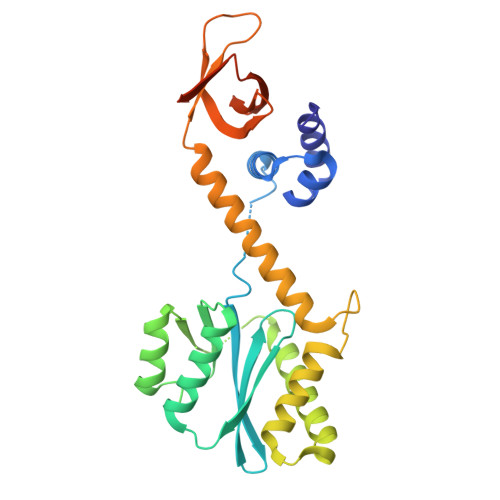
Integrase anchors viral RNA to the HIV-1 capsid interior.
Singer, M.R., Li, Z., Rey, J.S., Hope, J., Chenavier, F., Cook, N.J., Punch, E., Smith, J., Zhou, Z., Maslen, S., Masino, L., Nans, A., Skehel, M., Taylor, I.A., Zanetti, G., Zhang, P., Perilla, J.R., Engelman, A.N., Cherepanov, P.(2026) Nature 652: 1068-1075
- PubMed: 41708858 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-026-10154-x
- Primary Citation Related Structures:
9RMU, 9RMX - PubMed Abstract:
HIV-1 integrase (IN) promotes encapsulation of viral genomic RNA into mature viral cores, and this function is a target for ongoing antiretroviral drug development efforts 1-3 . Here we determined the cryogenic electron microscopy (cryo-EM) structure of a primate lentiviral IN in a complex with RNA, revealing a linear filament made of IN octamer repeat units, each comprising a pair of asymmetric homotetramers. The assembly is stabilized through IN-RNA interactions involving mainly the IN C-terminal domains and RNA backbone. The spacing and orientation of the IN filament repeat units closely matched those of consecutive capsid (CA) hexamers within the mature CA lattice. Using cryo-EM images of native purified HIV-1 cores, we refined the structure of the IN filament as it propagates along the luminal side of the CA lattice. Each IN tetramer within the filament nestled in a CA hexamer, engaging closely with the major homology regions. Substitutions of residues involved in IN-CA contacts yielded eccentric virions with RNA nucleoids located outside of the cores. Collectively, our results establish the structural basis for the HIV-1 IN-RNA interaction and reveal that IN forms an RNA-binding module on the luminal side of the mature CA lattice.
- Chromatin Structure & Mobile DNA Laboratory, The Francis Crick Institute, London, UK.
Organizational Affiliation: