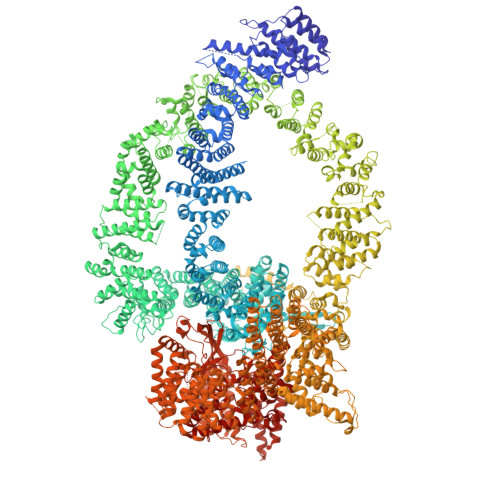
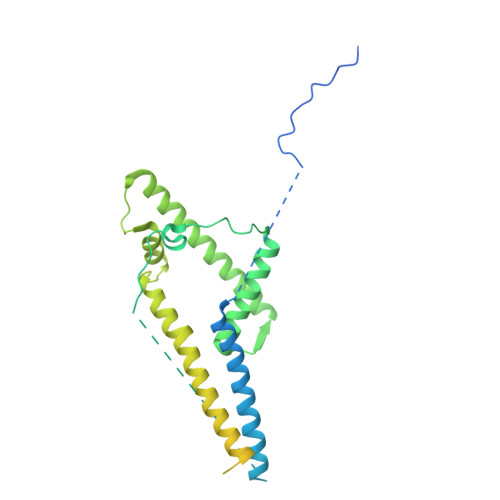
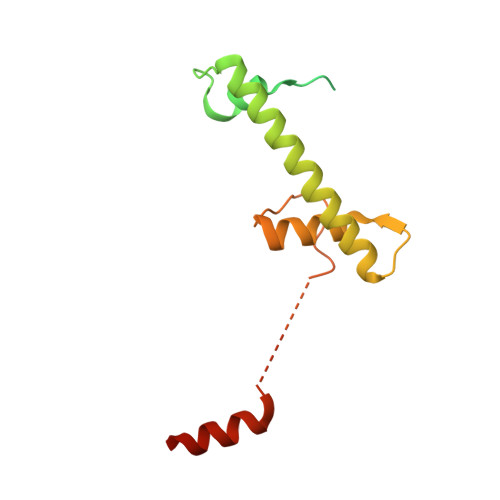
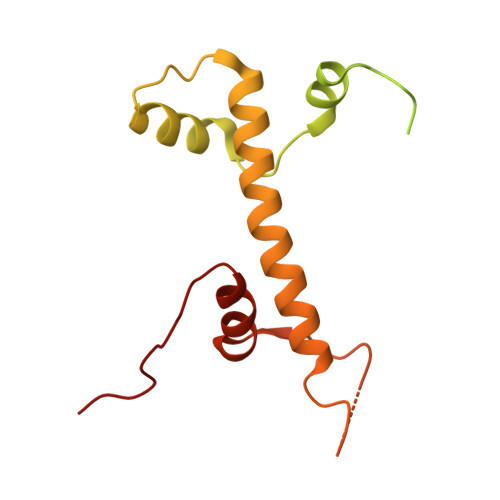
Insights into the structure and evolution of the human SAGA complex by affinity-ligand purification.
Damilot, M., Schoeps, T., Tora, L., Schultz, P., Lebeau, L., Papai, G., Ben-Shem, A.(2026) Sci Adv 12: eaec8104-eaec8104
- PubMed: 41849588 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.aec8104
- Primary Citation Related Structures:
9RDK - PubMed Abstract:
Human SAGA is a 20-subunit transcriptional coactivator. Compared with yeast, metazoan SAGA uniquely incorporates a 150-kDa splicing-factor module (SPL), also present in U2 small nuclear ribonucleoprotein (U2snRNP). Metazoan gene duplication further specialized shared TFIID/SAGA subunits into SAGA-specific paralogs (TAF5L and TAF6L), but the functional consequences of this divergence are unknown. We report the structure of endogenous human SAGA purified via an affinity ligand from cells that were not disturbed by any genomic engineering tools. Our work reveals the high-resolution structure of SPL and the TAF6L HEAT repeat domain that provides the SPL with a docking surface. We elucidate how SPL and the HEAT repeats are incorporated into SAGA. We identify major structural differences between TAF6L/TAF5L and their canonical paralogs that enable SPL accommodation. SPL engages SAGA through a substantially smaller interface than in U2snRNP, despite sharing a deeply inserted helical motif. The seemingly weaker interaction of SPL with SAGA raises the possibility that SAGA relays this module to the splicing machinery.
- Department of Integrated Structural Biology, Institut de Génétique et de Biologie Moléculaire et Cellulaire (IGBMC), Illkirch, France.
Organizational Affiliation: