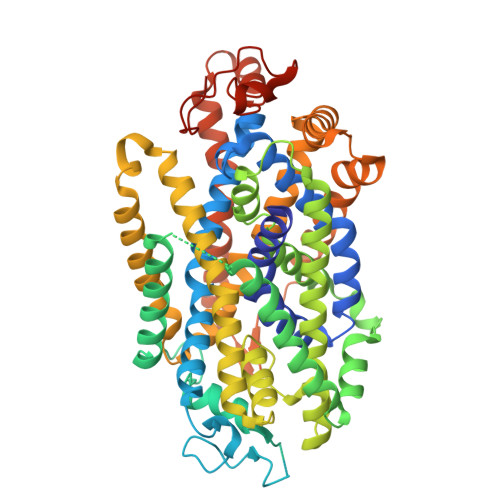
A reversible allosteric inhibitor of GlyT2 alleviates neuropathic pain without on-target side effects.
Cantwell Chater, R.P., Peiser-Oliver, J., Pati, T.K., Quinn, A.S., Lotsaris, I., Frangos, Z.J., Anderson, K.E., Tischer, A.E., Williams-Noonan, B.J., Aubrey, K.R., O'Mara, M.L., Michaelides, M., Mohammadi, S.A., Cioffi, C.L., Vandenberg, R.J., Shahsavar, A.(2025) bioRxiv
- PubMed: 41332641 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1101/2025.04.21.649698
- Primary Citation Related Structures:
9HUE, 9HUF, 9R1H - PubMed Abstract:
Chronic neuropathic pain, caused by nerve damage or disease, is increasing in prevalence, but current treatments are ineffective and over-reliant on opioids. The neuronal glycine transporter, GlyT2, regulates inhibitory glycinergic neurotransmission and represents a promising target for new analgesics. However, most GlyT2 inhibitors cause significant side effects, in part due to irreversible inhibition at analgesic doses. Here we develop a reversible inhibitor of GlyT2, RPI-GLYT2-82, and identify its binding site by determining cryo-EM structures of human GlyT2. We capture three fundamental conformational states of GlyT2 in the substrate-free state, and bound to either glycine, RPI-GLYT2-82 or the pseudo-irreversible inhibitor ORG25543. We demonstrate that RPI-GLYT2-82 dissociates from GlyT2 faster than ORG25543, providing analgesia in mouse neuropathic pain models without on-target side-effects or addiction liability. Our data provide a mechanistic understanding of allosteric inhibition of glycine transport, enabling structure-based design of non-opioid analgesics.
- Department of Drug Design and Pharmacology, Faculty of Health and Medical Sciences, University of Copenhagen, Copenhagen, Denmark.
Organizational Affiliation: