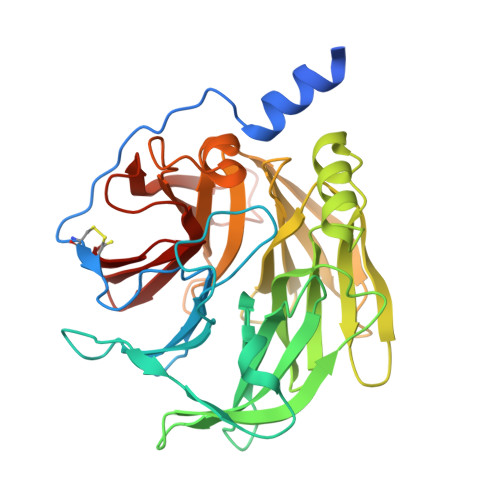
Intramolecular sensitization and structure of a Tb 3+ /2-hydroxyquinoline conjugate in the paraoxonase 1 active site.
Smerkolj, J., Bahun, M., Poklar Ulrih, N., Bavec, A., Pavsic, M., Golicnik, M.(2025) Dalton Trans 54: 12471-12481
- PubMed: 40735812 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1039/d5dt01484k
- Primary Citation Related Structures:
9R0Q - PubMed Abstract:
Paraoxonase 1 (PON1) is a Ca 2+ -dependent enzyme involved in oxidative stress processes and is widely studied for its protective roles in various diseases. Intermolecular sensitization of lanthanide ions was implemented by replacing Ca 2+ ions from the recombinant PON1 (rePON1) catalytic site in the presence of 2-hydroxyquinoline (2HQ) as an external antenna. Although the replacement of Ca 2+ ions with lanthanide ions indicates weaker binding affinity for the coordination of 2HQ in the protein milieu of the rePON1 active site, it results in the formation of a highly emissive supramolecular complex in the case of Tb 3+ ions. The architecture of the ternary rePON1 : Tb 3+ : 2HQ conjugate, which allows efficient terbium sensitization and its specific long-wavelength metal phosphorescence emission, was resolved by X-ray crystallography. These findings could establish a non-catalytic quantification strategy for PON1 and provide additional structural insights into lanthanide substitution in this Ca 2+ -dependent enzyme.
- University of Ljubljana, Faculty of Medicine, Institute of Biochemistry and Molecular Genetics, Vrazov trg 2, 1000 Ljubljana, Slovenia. marko.golicnik@mf.uni-lj.si.
Organizational Affiliation: