N-terphenylpicolinamide derivatives designed to target PD-L1 increase activation and proliferation of T cells, and their cytotoxic properties toward cancer cells.
Muszak, D., Kocik-Krol, J., Zaber, J., Kruc, O., Palej, U., Fijolkowska, K., Maslanka, A., Magiera-Mularz, K., Plewka, J., Stec, M., Surmiak, M., Szafarz, M., Siedlar, M., Musielak, B., Kitel, R., Wyska, E., Skalniak, L., Surmiak, E.(2026) Eur J Med Chem 307: 118652-118652
- PubMed: 41691995
- DOI: https://doi.org/10.1016/j.ejmech.2026.118652
- Primary Citation of Related Structures:
9QSM - PubMed Abstract:
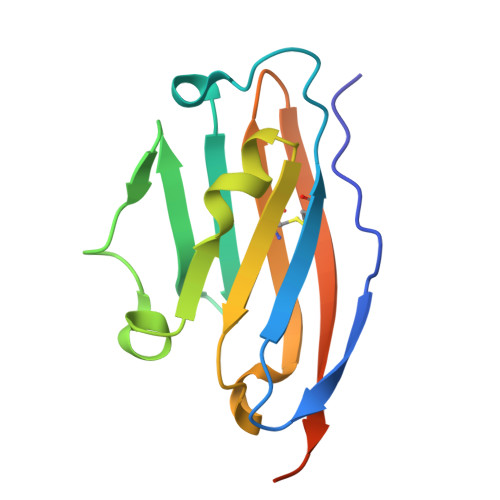
Programmed Cell Death Protein-1 (PD-1)/Programmed Cell Death-Ligand 1 (PD-L1) interaction has a crucial role in maintaining the immune system's self-tolerance by downregulating T cell activation. This mechanism is also used by several types of cancers. By overexpressing the PD-L1 protein, cancer cells can evade the immune response and, therefore, become invisible to the immune system. Herein, we present a detailed characterization of the activity of improved N-terphenylpicolinamides, a class of small molecular blockers targeting the PD-L1 protein disclosed in our recent patent and following patent applications. In our studies, we utilized a cell-based structure-activity relationship (SAR) analysis, which allowed us to discriminate the bioactivity of molecules beyond the detection limits of the protein-based HTRF assay. Our final molecules display high affinity to the molecular target and in vitro bioactivity approaching the activity of a positive control ARB-272572 molecule. An optimized molecule activates primary immune cells, leading to enhanced elimination of cancer cells, as we show in a newly developed co-culture setup. In addition, a co-crystal structure described here confirms the intended mode of binding of the small molecule to PD-L1. Our pharmacokinetics (PK) results rationalize the choice of a representative molecule for further in vivo testing.
- Jagiellonian University, Faculty of Chemistry, Department of Organic Chemistry, Gronostajowa 2, Krakow, 30-387, Poland. Electronic address: damian.muszak@uj.edu.pl.
Organizational Affiliation: