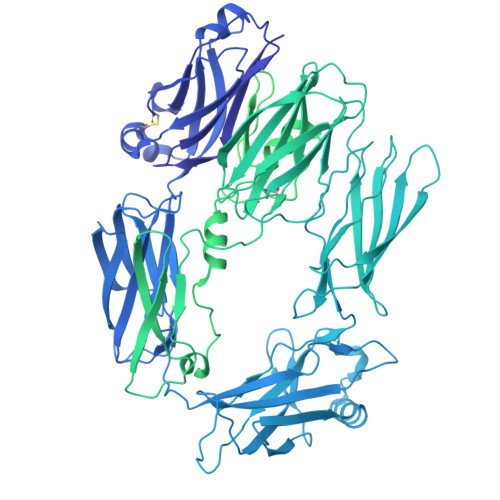
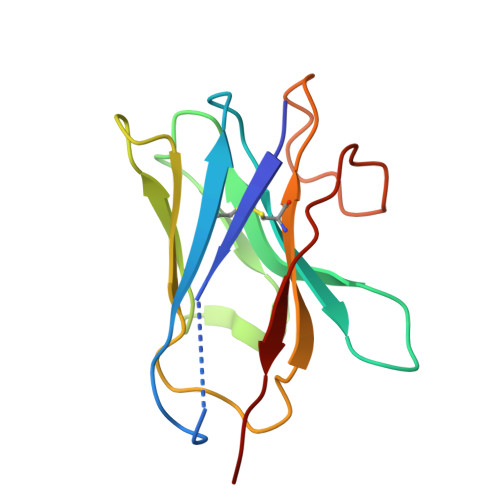
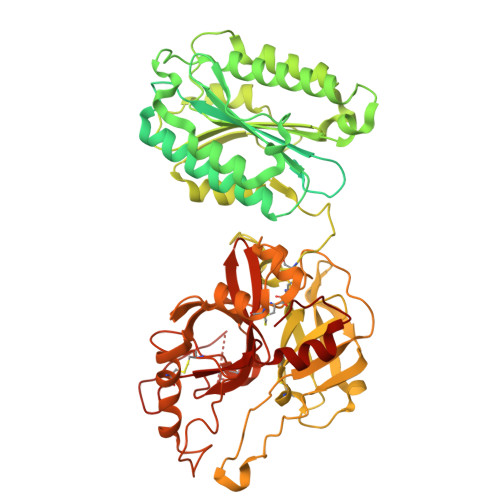
Structural insights into C3 convertase activity of the classical pathway of complement.
De la O Becerra, K.I., Brondijk, T.H.C., Serna Martin, I., Gros, P.(2025) Nat Commun 17: 993-993
- PubMed: 41413058 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-67730-4
- Primary Citation Related Structures:
9QJ4, 9QJ5, 9QK2, 9QPY - PubMed Abstract:
Immune protection by the complement system depends on C3 cleavage by C3 convertases that is critical to all three activation pathways. Structural data on convertase formation in the classical pathway and on C3-substrate binding to convertases is lacking. We present the cryo-EM structures of the proconvertase (C4b2), convertase (C4b2b), and convertase-substrate complex (C4b2b-C3) of the classical pathway. The data show that C2 and C4b form proconvertases and convertases like factor B and C3b of the alternative pathway. Substrate C3 binds C4b of the convertase through two interfaces: one also found in the SCIN-inhibited C3bBb dimer, and another facilitated by conformational changes in C3. Bending of C3 and swinging of the C2 protease bring the C3-scissile loop into the active site. The second, charged, C4b-interaction site favors C3- substrate binding, but upon cleavage repels product C3b. Thus, a charge switch-over mechanism effects the catalytic turnover of the convertases producing opsonin C3b.
- Structural Biochemistry, Bijvoet Centre for Biomolecular Research, Dept. of Chemistry, Faculty of Science, Utrecht University, Utrecht, The Netherlands.
Organizational Affiliation: