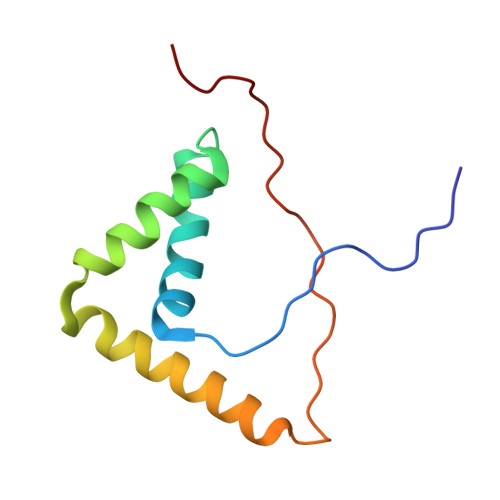
Solution structure of the Sox2 DNA-binding domain reveals conformational selection in DNA binding.
Orsetti, A., Slejfer, J., Ha, S., Kevelam, D.I., Tekkelenburg, J., van Duijn, T., Leppakoski, A., Sedrakyan, A., Szilagyi, A., Schellevis, R.D., Soufi, A., Cojocaru, V., van Ingen, H.(2025) Nucleic Acids Res 53
- PubMed: 41189057
- DOI: https://doi.org/10.1093/nar/gkaf1121
- Primary Citation Related Structures:
9QPF - PubMed Abstract:
The transcription factor Sox2 is a master regulator of cell pluripotency. While structural studies have provided insights into its DNA-bound conformation, the mechanisms governing its free-state conformational dynamics and DNA recognition remain elusive. Based on solution NMR spectroscopy and supported by molecular dynamics simulations, we here report the solution structure of the Sox2 DNA-binding domain (DBD), revealing that its helical core is well-structured and arranged as in its DNA-bound state. The folded, free protein coexists in dynamic equilibrium with partially unfolded states, which are quenched upon specific DNA binding. We show that the electrostatic environment significantly influences the Sox2-DBD stability, with high ionic strength stabilizing the protein. NMR titration experiments demonstrate that the nonspecific and specific DNA binding interfaces of Sox2 largely overlap. Specific binding, however, uniquely involves rigidification of part of the C-terminal tail. Based on these findings, we propose that the helical core of the Sox2-DBD is stabilized in its DNA-bound form prior to binding. Binding of Sox2 to DNA thus involves conformational selection, rather than exclusively induced fit, as was previously proposed. Through its pre-folded, DNA-binding competent fold, Sox2 may be able to rapidly switch from scanning of DNA to specific binding of its cognate site.
- NMR Spectroscopy Group, Bijvoet Centre for Biomolecular Research, Utrecht University, Utrecht, CH 3854, The Netherlands.
Organizational Affiliation: