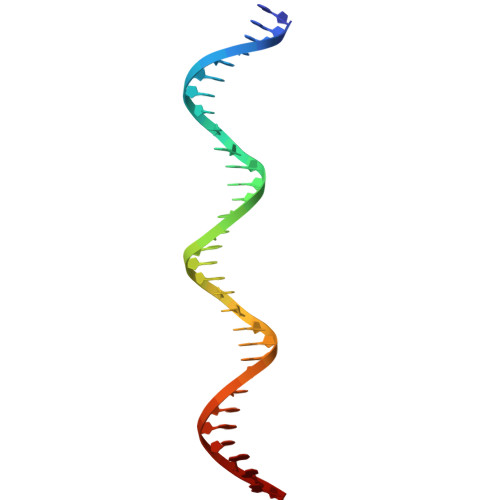
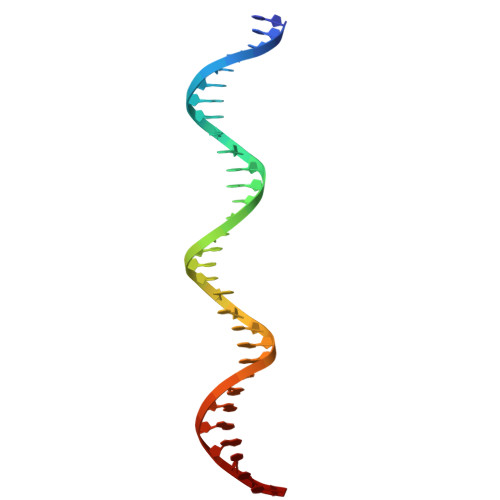
Structural basis of Nuclear Factor 1-X DNA recognition provides prototypic insight into the NFI family.
Tiberi, M., Lapi, M., Gourlay, L.J., Chaves-Sanjuan, A., Polentarutti, M., Demitri, N., Cavinato, M., Bonnet, D.M.V., Taglietti, V., Righetti, A., Sala, R., Cauteruccio, S., Kumawat, A., Russo, R., Barbiroli, A.G., Gnesutta, N., Camilloni, C., Bolognesi, M., Messina, G., Nardini, M.(2025) Nat Commun 16: 10170-10170
- PubMed: 41261119 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-65186-0
- Primary Citation Related Structures:
7QQD, 9QKY - PubMed Abstract:
Nuclear Factor I (NFI) proteins are involved in adenovirus DNA replication and regulate gene transcription, stem cell proliferation, and differentiation. They play key roles in development, cancer, and congenital disorders. Within the NFI family, NFI-X is critical for neural stem cell biology, hematopoiesis, muscle development, muscular dystrophies, and oncogenesis. Here, we present the structural characterization of the NFI transcription factor NFI-X, both alone and bound to its consensus palindromic DNA site. Our analyses reveal a MH1-like fold within NFI-X DNA-binding domain (DBD) and identify crucial structural determinants for activity, such as a Zn²⁺ binding site, dimeric assembly, and DNA-binding specificity. Given the ~85% sequence identity within the NFI DBDs, our structural data are prototypic for the entire family, a NFI Rosetta Stone that allows decoding a wealth of biochemical and functional data and provides a precise target for drug design in a wider disease context.
- Department of Biosciences, Università degli Studi di Milano, Milano, Italy.
Organizational Affiliation: