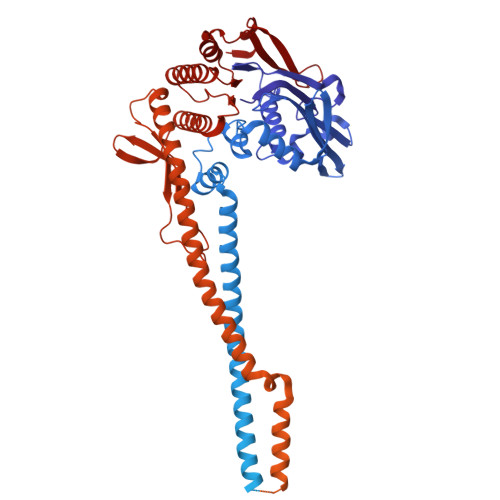
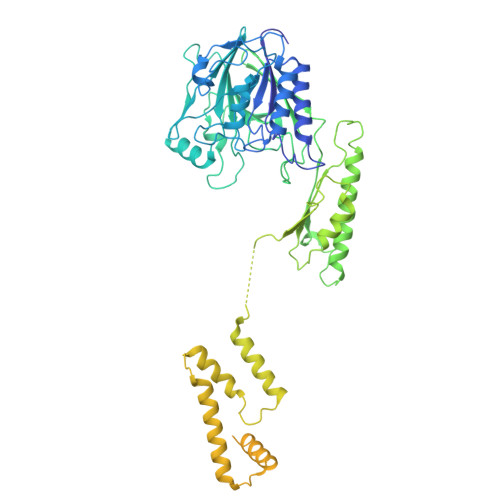
Structural basis for DNA break sensing by human MRE11-RAD50-NBS1 and its regulation by telomeric factor TRF2.
Fan, Y., Kuybu, F., Cui, H., Lammens, K., Chen, J.X., Kugler, M., Jung, C., Hopfner, K.P.(2025) Nat Commun 16: 8320-8320
- PubMed: 40968163 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-64082-x
- Primary Citation Related Structures:
9Q9H, 9Q9I, 9Q9J, 9Q9K, 9Q9M - PubMed Abstract:
The MRE11-RAD50-NBS1 (MRN) complex is a central, multifunctional factor in the detection, signaling and nucleolytic processing of DNA double-strand breaks (DSBs). To clarify how human MRN binds generic and telomeric DNA ends and can separate DNA end sensing from nuclease activities, we determined cryo-electron microscopy (cryo-EM) structures of human MRN bound to DNA and to DNA and the telomere protection factor TRF2. MRN senses DSBs through a tight clamp-like sensing state with closed coiled-coil domains, but auto-inhibited MRE11 nuclease. NBS1 wraps around the MRE11 dimer, with NBS1's ATM recruitment motif sequestered by binding to the regulatory RAD50 S site, necessitating a switch in the NBS1 C helix for ATM activation. At telomeric DNA, TRF2 blocks the second S site via the iDDR motif to prevent nuclease and ATM activation. Our results provide a structural framework for DNA sensing via a gating mechanism and separation of sensing, signaling and processing activities of mammalian MRN.
- Gene Center, Department of Biochemistry, Ludwig-Maximilians-Universität München, Feodor Lynen Straße 25, 81377, Munich, Germany.
Organizational Affiliation: