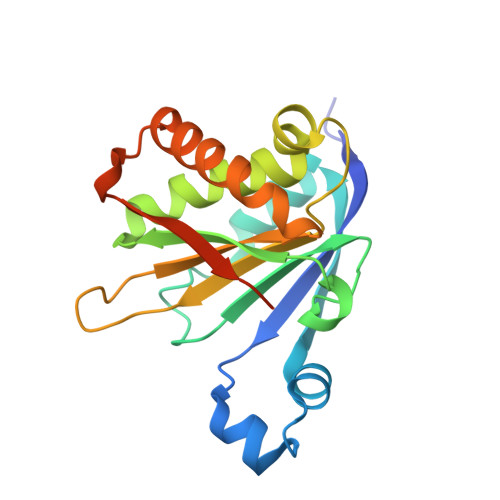
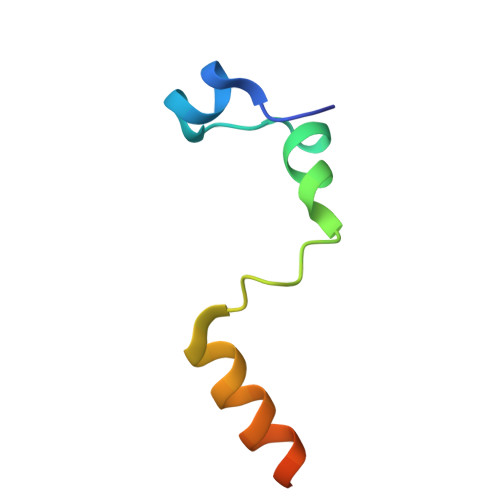
Selective eIF4E-eIF4G Pairing and Cap-4 Recognition Mechanisms in Trypanosomatids: Insights From EIF4E5-EIF4G1 and EIF4E6-EIF4G5 Complexes.
Penteado, R.F., Vichier-Guerre, S., da Silva Pereira, B.M., Dugue, L., Guerra Slompo, E.P., Assuncao de Matos, T.R., Pochet, S., Zanchin, N.I.T., Guimaraes, B.G.(2026) J Mol Biology 438: 169550-169550
- PubMed: 41265746 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2025.169550
- Primary Citation Related Structures:
9Q72, 9Q73, 9Q74, 9Q75, 9Q78, 9Q79 - PubMed Abstract:
Translation initiation in eukaryotes is a highly regulated process involving the assembly of several transient protein complexes. A key step in this process is recognition of the mRNA 5' cap structure by the initiation factor eIF4E, a core component of the eIF4F complex. In trypanosomatids, this mechanism diverges from canonical eukaryotic systems, featuring five distinct eIF4F-like complexes formed through specific pairings of eIF4E and eIF4G homologs. Additionally, trypanosomatid mRNAs exhibit a unique hypermethylated cap-4 structure at the 5' end. To elucidate the molecular basis of selective eIF4E-eIF4G interactions and the modulation of cap binding upon eIF4G engagement, we determined high-resolution crystal structures of EIF4E5-EIF4G1 complexes from Trypanosoma brucei and T. cruzi, and the EIF4E6-EIF4G5 complex from T. cruzi. These structural studies, supported by biophysical analyses in the presence and absence of a cap-4 analog, reveal key determinants of cap recognition associated with cap-4 structural flexibility and plasticity in the cap-binding pockets. We observe conformational rearrangements upon eIF4G binding and propose a relationship between these structural changes and increased cap-4 affinity. In addition, comparative structural analysis of the EIF4E5-EIF4G1 and EIF4E6-EIF4G5 complexes offers atomic-level insights into the molecular determinants of specificity that govern selective eIF4E-eIF4G pairings in trypanosomatids.
- Carlos Chagas Institute, Oswaldo Cruz Foundation, Rua Prof. Algacyr Munhoz Mader, 3775, 81350-010 Curitiba, PR, Brazil.
Organizational Affiliation: