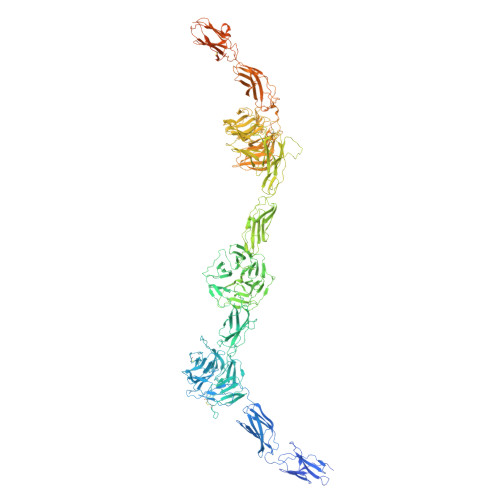
Clustering and a conformational switch drive activation of the mammalian receptor tyrosine kinase ROS1.
Li, H., Zhang, J., Li, T., Wang, Y., Alarcon, C.R., Klein, D.E.(2026) Nat Commun 17
- PubMed: 41698918 Search on PubMed
- DOI: https://doi.org/10.1038/s41467-026-69630-7
- Primary Citation Related Structures:
10FT, 10GH, 9DZ4, 9PVP, 9PWQ - PubMed Abstract:
Receptor tyrosine kinases (RTKs) are key regulators of cellular signaling and are often co-opted in cancer. ROS1 is an orphan RTK aberrantly expressed in multiple tumors, yet no approved biologic therapies target it, and its activation mechanism remains unknown. Here, we present Cryo-EM structures of mammalian ROS1 in ligand-free and NELL2-bound states, revealing how trimeric NELL2 induces both receptor clustering and a conformational switch that relieves receptor autoinhibition - both mechanisms are required for ROS1 activation. These structures, along with biochemical characterization, reflect a striking evolutionary divergence in regulatory logic compared to the invertebrate ortholog Sevenless (dROS1), highlighting how conserved RTKs can adopt fundamentally different activation strategies. Guided by these structural insights, we develop monoclonal antibodies that either block ligand binding or trap ROS1 in an inactive conformation. These agents potently suppress ROS1 signaling, representing distinct mechanistic classes of biologics that directly target ROS1 activity. Our findings elucidate a distinct mode of RTK regulation and establish a therapeutic framework for cancers driven by ROS1.
- Department of Pharmacology, Yale School of Medicine, New Haven, CT, USA.
Organizational Affiliation: