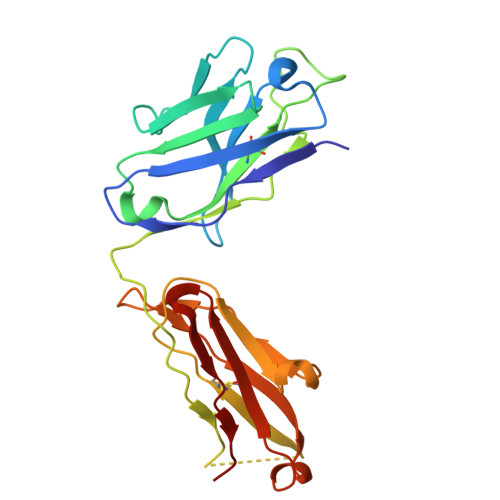
Affinity maturation and light-chain-mediated paratope diversification anticipates viral evolution.
Dingus, J., Yoo, D.K., Kumar, S., Wang, Y., Kibria, M.G., Saghaei, S., Allahyari, Z., Chen, J.W., Caputo, N.M., Hwang, J., Chen, B., Wesemann, D.R.(2025) Cell Rep 45: 116800-116800
- PubMed: 41474620 Search on PubMed
- DOI: https://doi.org/10.1016/j.celrep.2025.116800
- Primary Citation Related Structures:
9PW4 - PubMed Abstract:
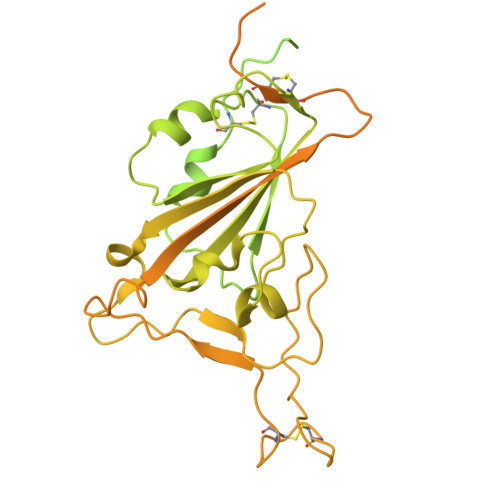
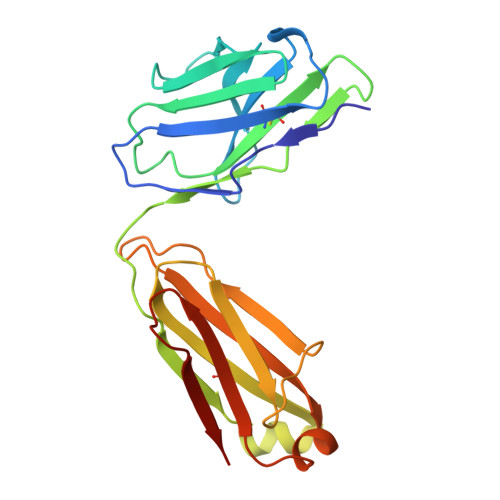
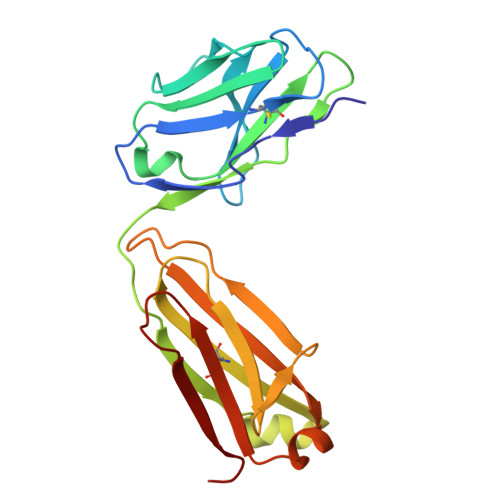
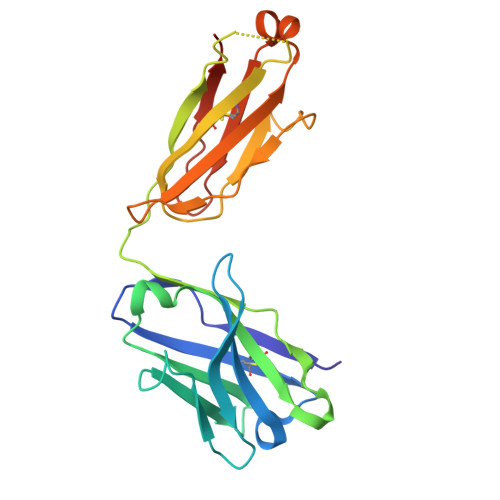
A key goal of vaccinology is to train the immune system to combat current pathogens while preparing it for future variants. Here, we investigate how Wuhan strain severe acute respiratory syndrome coronavirus 2 mRNA vaccination generates "anticipatory breadth" in an antibody family exhibiting germline complementarity to the ACE2 binding site on the receptor-binding-domain (RBD). IGHV3-53/66 antibodies from infection-naive vaccinees frequently neutralize Omicron variants and contain hallmark breadth-enhancing mutations. While Omicron breakthrough infection does not alter IGHV3-53/66 mutation frequencies, it modifies Ig light-chain pairing frequencies, suggesting variant-driven selection for favorable pairings. Structural analyses of IGHV3-53/66-RBD complexes show that hallmark heavy-chain mutations refine interactions with conserved RBD residues, while alternative Ig light-chain pairings modify contacts at Omicron mutation sites. Together, these findings support a cooperative model of anticipatory breadth involving targeting of a functionally constrained epitope, affinity maturation to establish an affinity buffer, and alternative Ig light-chain pairings to diversify paratopes-providing a mechanistic framework for anticipating viral evolution.
- Department of Medicine, Division of Allergy and Clinical Immunology, Division of Genetics, Brigham and Women's Hospital, Harvard Medical School, Boston, MA 02115, USA; Broad Institute of MIT, and Harvard, Cambridge, MA 02139, USA; Ragon Institute of MGH, MIT, and Harvard, Cambridge, MA 02139, USA.
Organizational Affiliation: