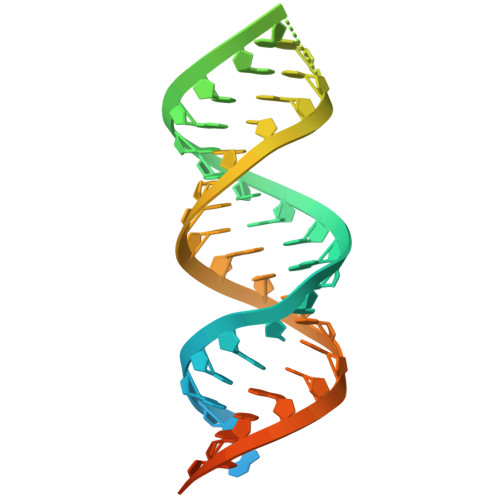
Structural characterization of the HDV virion and its ribonucleoprotein.
Itskanov, S., Ary, B., Mehra, U., Lew, I., Novikov, N., Schmitz, U., Holdorf, M.M., Beran, R.K., Lansdon, E.B.(2026) Proc Natl Acad Sci U S A 123: e2519809123-e2519809123
- PubMed: 41564123 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2519809123
- Primary Citation Related Structures:
9PRC - PubMed Abstract:
Hepatitis D virus (HDV) is a small RNA satellite virus of hepatitis B virus (HBV) which encodes a single protein, HDV delta antigen (HDAg), that is required for replication. Viral replication occurs independently from HBV and relies primarily on host RNA polymerase(s). Bulevirtide, a viral entry inhibitor, is the only approved treatment for chronic HDV but has a low cure rate as a monotherapy, and most patients rebound following cessation of therapy. It is likely that an inhibitor targeting HDV replication is necessary to achieve HDV cure, but the paucity of HDV-derived elements and limited understanding of HDV replication presents a significant therapeutic challenge. Understanding the precise mechanism of interactions between HDAg and viral RNA, and how it is packaged within the virion can inspire structure-guided drug design targeting replication. Using cryoelectron tomography and single particle cryoelectron microscopy, we present reconstructions of the virion and viral RNPs. We observed multiple binding configurations in vitro that suggest a propensity to arrange four RNA segments around repeating units of HDAg in a ladder-like formation. The oligomerization domains of a homo-octameric HDAg complex are directly involved in RNA binding by utilizing the vertices and sides of its square-shaped architecture to bind RNA in a sequence-promiscuous fashion. Structure-function analysis reveals that these RNA contact sites are important for viral replication and their disruption may be a potential avenue for next-generation antivirals to treat HDV.
- Structural Biology & Chemistry, Gilead Sciences, Inc., Foster City, CA 94404.
Organizational Affiliation: