Development of a PRAME pMHC targeted T cell engager for solid tumor therapy.
Skrzypczynska, K., Schimert, K., Stephenson, H., Mah, I.K., Mortenson, D., Boyd, K., Hardman, T., Novikov, N., Seto, E., Lu, S., Yen, R., Lee, B., Wang, M., Kang, D., Huang, Y., Yu, X., Hung, M., Ding, S., Thomsen, N., Oakdale, N.S.(2025) MAbs 17: 2563773-2563773
- PubMed: 40994043 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1080/19420862.2025.2563773
- Primary Citation Related Structures:
9PKV - PubMed Abstract:
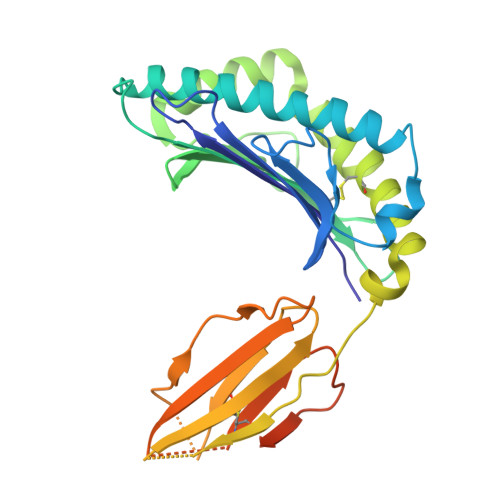
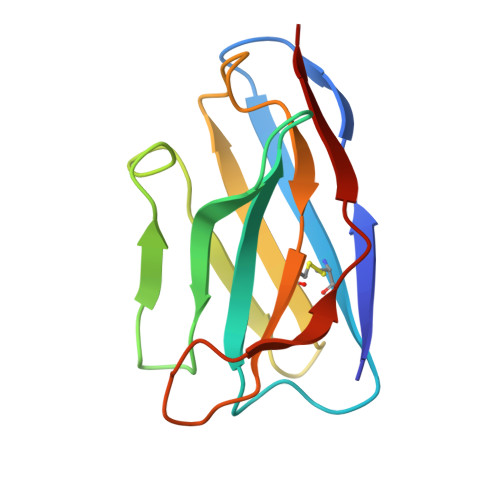
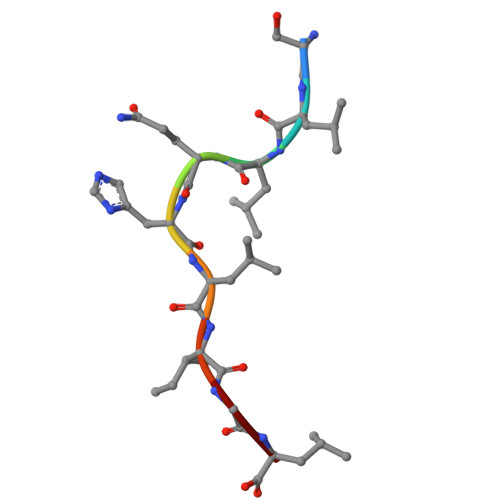
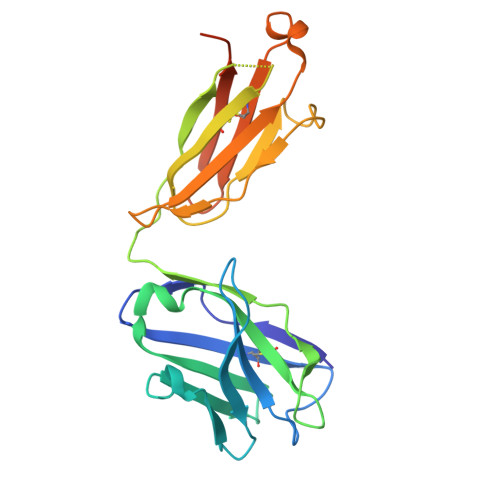
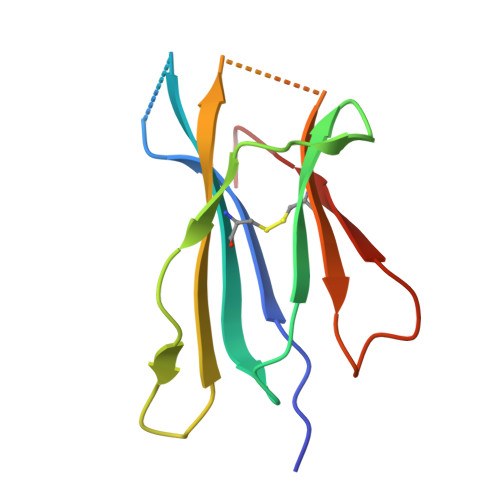
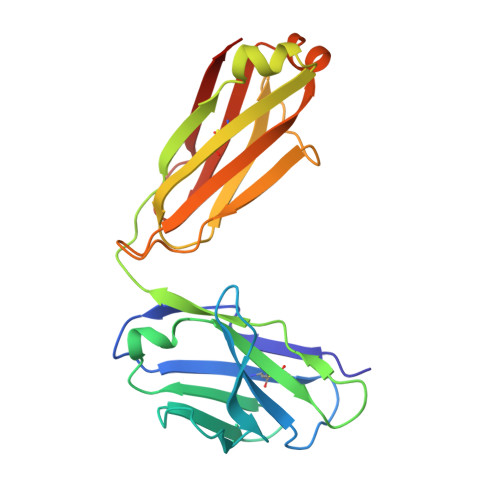
Bispecific T cell engager (TCE) therapies have demonstrated transformative clinical success in the treatment of hematological cancers, but the lack of antigens that are sufficiently selective for malignant cells has hampered the success of TCEs in the solid-tumor space. To overcome the on-target, off-tumor toxicities that result from the expression of even low levels of tumor-associated antigens in healthy tissues, we sought to identify a TCE target with highly tumor-restricted expression patterns. Here, we characterize cancer-testes antigen Preferentially Expressed Antigen in Melanoma (PRAME) as a highly selective tumor antigen and identify a proteasomal degradation peptide PRAME 425-433 (PRAME 425 ) presented in the context of major histocompatibility complex I (MHCI) as an attractive TCE target. We designed a TCR-mimic (TCRm) antibody screening cascade that prioritizes screening anti-PRAME pMHC binders in off-target T cell dependent cellular cytotoxicity assays in a potent TCE format, rather than relying solely on traditional pMHC binding assays, to determine specificity. Using this screening cascade, we discovered antibodies that selectively bind PRAME 425 pMHC without over-recognition of off-target peptides or MHCI via a TCR-like binding geometry. We further solved the first structure of an anti-PRAME 425 pMHC TCRm antibody in complex with PRAME 425 /HLA-A *02:01 using cryo electron microscopy to confirm the TCRm antibody binds in a TCR-like binding geometry and specifically recognizes the PRAME 425 peptide. By formatting these novel TCRm antibodies into potent TCEs, we demonstrate PRAME 425 pMHC-specific killing of tumor cells, representing a new class of anti-PRAME pMHC biologics.
- Oncology, Gilead Sciences, Inc, Foster City, CA, USA.
Organizational Affiliation: