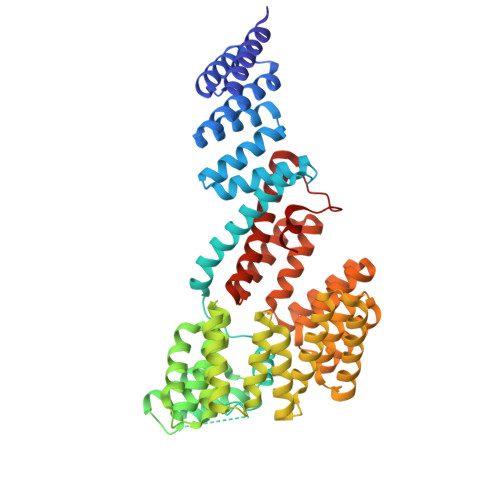
An allosteric network governs Tom70 conformational dynamics to coordinate mitochondrial import.
Bachochin, M.J., McGuire, K.L., Cook, B.D., Ye, Q., Silletti, S., Corbett, K.D., Komives, E.A., Herzik Jr., M.A.(2026) Structure 34: 273
- PubMed: 41386227
- DOI: https://doi.org/10.1016/j.str.2025.11.011
- Primary Citation Related Structures:
9PKQ - PubMed Abstract:
Tom70 mediates mitochondrial protein import by coordinating transfer of cytosolic preproteins from Hsp70/Hsp90 to the translocase of the outer membrane (TOM) complex. In humans, the cytosolic domain of Tom70 (HsTom70c) is entirely α-helical and comprises modular TPR motifs divided into an N-terminal chaperone-binding and a C-terminal preprotein-binding domain. However, the mechanisms linking these functional regions remain poorly understood. Here, we present the 2.04 Å crystal structure of unliganded HsTom70c, revealing two distinct conformations-open and closed-within the asymmetric unit. These states are stabilized by interdomain crystal contacts and supported in solution by hydrogen-deuterium exchange mass spectrometry (HDX-MS) and molecular dynamics (MD) simulations. Principal component and network analyses reveal a continuum of motion linking the NTD and CTD via key residues in helices α7, α8, and α25. Engagement of the CTD by viral protein Orf9b disrupts this network, stabilizing a partially closed intermediate and dampening distal NTD dynamics.
- Department of Chemistry and Biochemistry, University of California, San Diego, La Jolla, CA 92093, USA.
Organizational Affiliation: