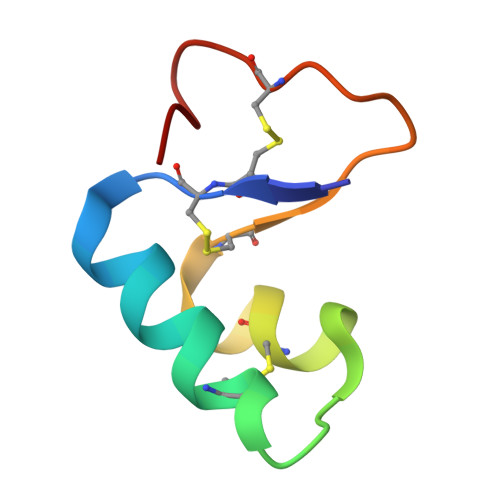
Direct from the seed: an atomic resolution protein structure by ab initio MicroED.
Vasireddy, P.C.R., Low-Beer, T., Spoth, K.A., Acehan, D., Crawley, M.R., Martynowycz, M.W.(2026) Nat Commun 17
- PubMed: 41690942 Search on PubMed
- DOI: https://doi.org/10.1038/s41467-026-69601-y
- Primary Citation Related Structures:
9PIS - PubMed Abstract:
While purifying the seed protein crambin, we find that needles of pure protein nanocrystals form spontaneously during the drying of a simple ethanolic purification drop. These needles diffract X-rays weakly but are well-suited for microcrystal electron diffraction (MicroED). Merging data from 58 such nanocrystals yields diffraction to 0.85 Å resolution and solves the structure ab initio using a five-residue helical fragment to initiate density modification. The resulting map enables automated model building and resolves individual hydrogen atoms. This work reports an atomic resolution MicroED structure of the protein crambin solved ab initio. This study establishes a publicly available benchmark showing that sub-ångström ab initio MicroED can be achieved on standard 200 kV instrumentation without energy filtering or FIB milling, when serial merging is combined with anisotropy aware truncation to address preferred orientation. For this dataset, isotropic truncation alone did not produce a fragment placement suitable for phasing in this workflow, whereas applying anisotropy correction supported an ab initio solution.
- Department of Structural Biology, Jacobs School of Medicine and Biomedical Sciences, University at Buffalo, The State University of New York, Buffalo, NY, USA.
Organizational Affiliation: