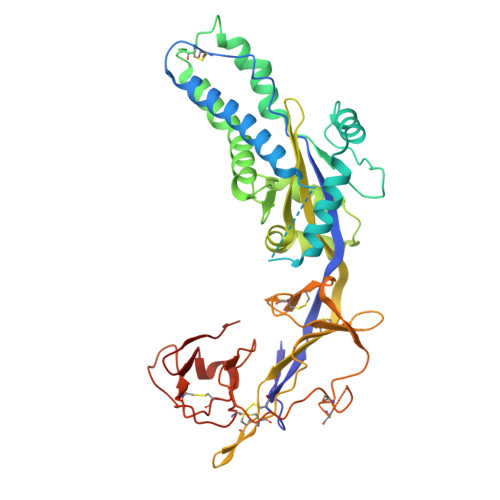
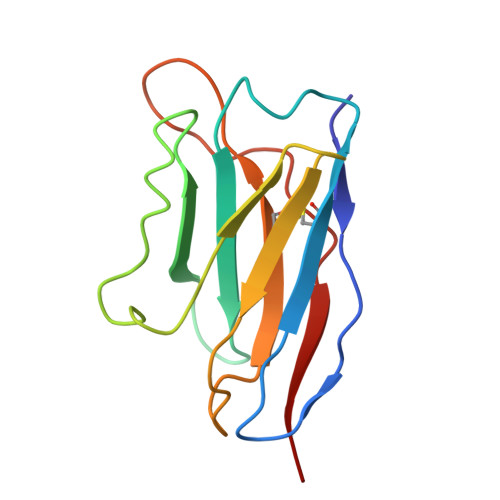
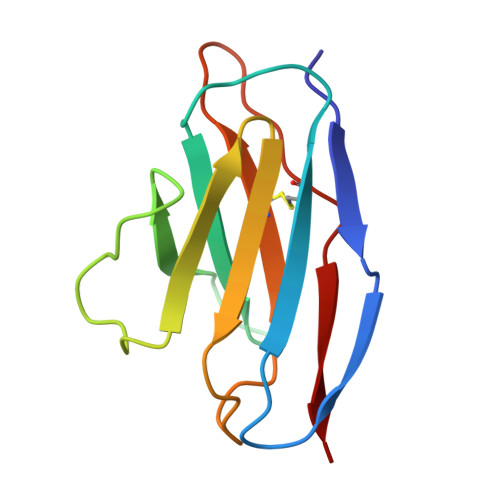
Structural basis for childhood antibody recognition of the human metapneumovirus fusion protein.
Khalil, A.M., Ghazi Esfahani, B., Miller, R.J., Huang, J., Kuan, G., Balmaseda, A., Gordon, A., Mousa, J.J.(2025) Nat Commun 17: 1267-1267
- PubMed: 41455691 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-68021-8
- Primary Citation Related Structures:
9OS5, 9PDX, 9PDY - PubMed Abstract:
Human metapneumovirus (hMPV) is a significant cause of acute respiratory illness in children and older adults, with most children becoming seropositive by five years of age. The hMPV fusion (F) protein is the sole target of neutralizing antibodies, and while most common B-cell-targeted neutralizing epitopes on the hMPV F protein have been determined in hMPV-infected adults, the antibody response in hMPV-infected children remains undefined. We isolate five human monoclonal antibodies (mAbs) from hMPV-infected children, and evaluate their binding avidity, neutralization potency, epitope specificity, and in vivo efficacy. All mAbs are neutralizing, and epitope binning reveals four different epitopes targeted by the mAbs. Cryo-EM structures of four mAbs in complex with the hMPV F protein reveal epitopes on the hMPV F trimer surface as well as an intratrimer epitope located completely within the hMPV F trimer interface. Furthermore, we determine the prophylactic efficacy of the mAbs in protection against hMPV challenge in mice. These findings provide new insights into the immunodominant antigenic epitopes on the hMPV F protein in children and identify new mAbs for hMPV F disease prevention.
- Department of Biomedical Sciences, College of Medicine, Florida State University, Tallahassee, FL, USA.
Organizational Affiliation: