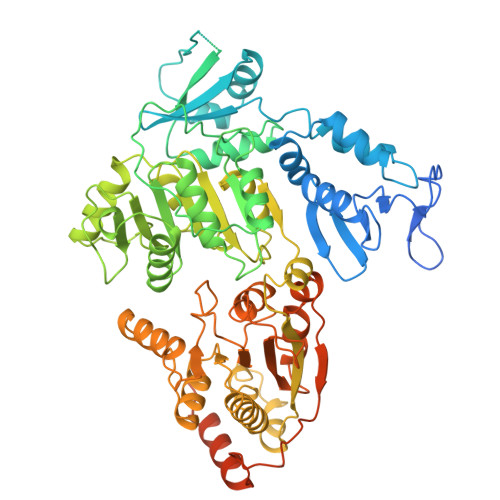
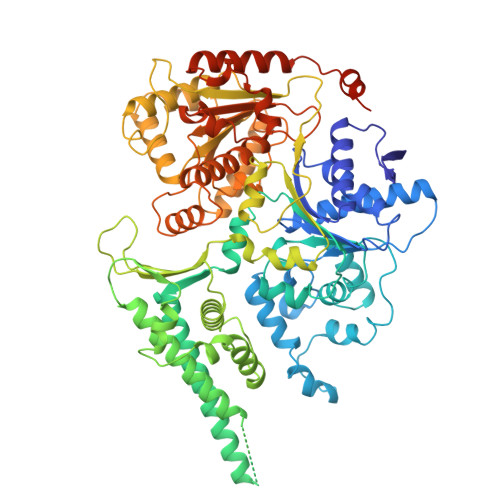
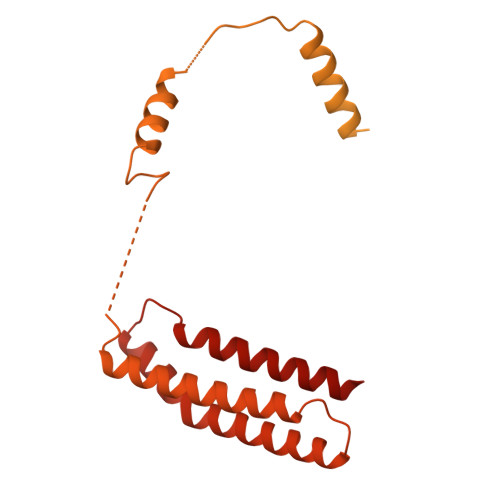
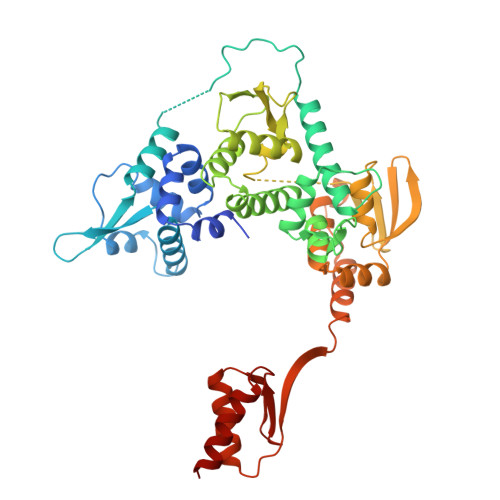
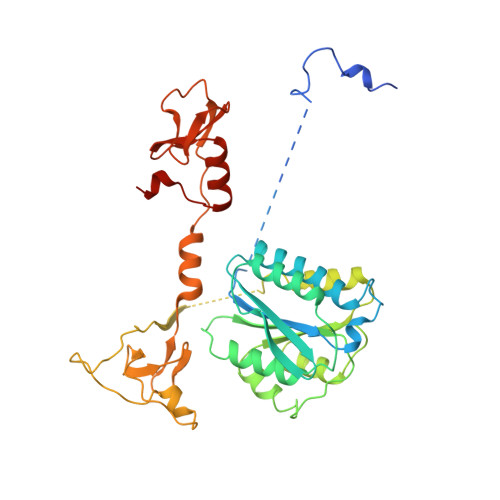
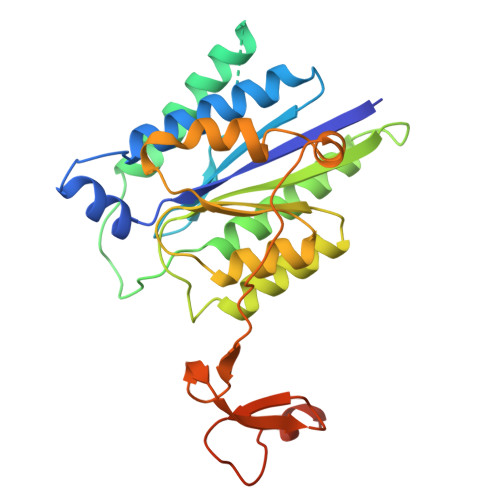
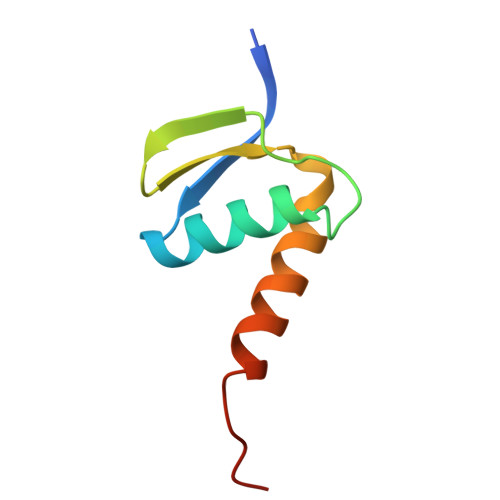
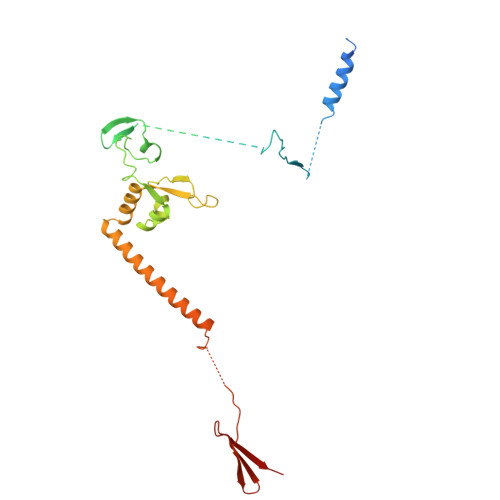
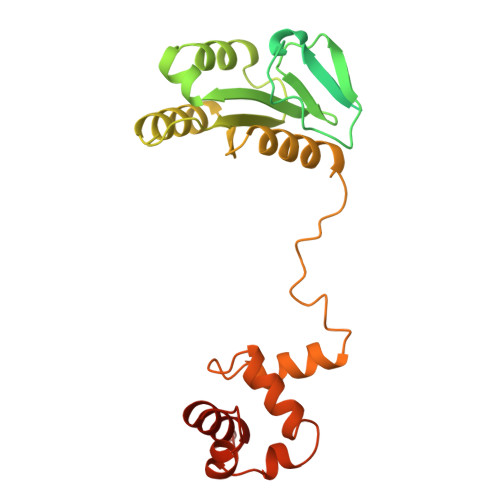
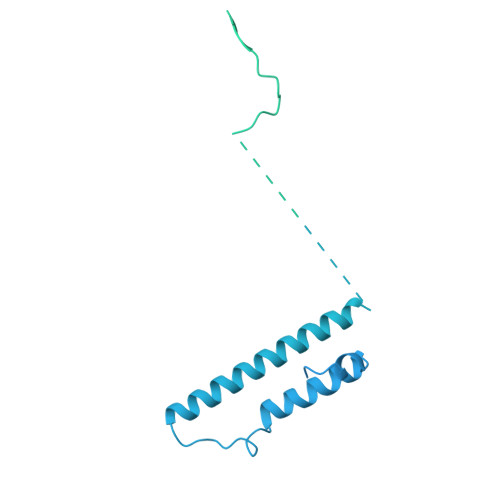
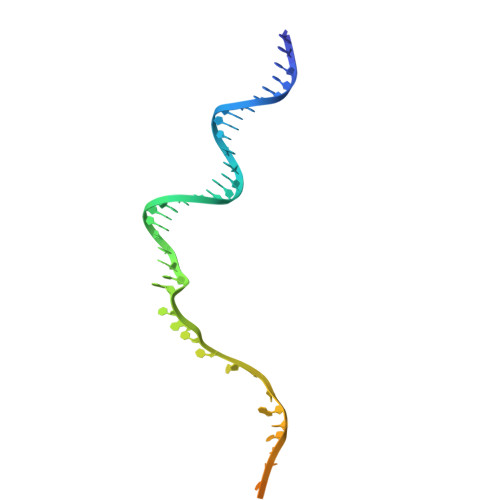
Pre-incision structures reveal principles of DNA nucleotide excision repair.
Li, E.C.L., Kim, J., Brussee, S.J., Sugasawa, K., Luijsterburg, M.S., Yang, W.(2026) Nature 652: 1060-1067
- PubMed: 41673165 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41586-026-10122-5
- Primary Citation Related Structures:
9PCP, 9PD3, 9PD4, 9PD5, 9XYU - PubMed Abstract:
Nucleotide excision repair (NER) removes bulky adducts from genomic DNA and prevents the ultraviolet light-sensitivity disease xeroderma pigmentosum, cancer and premature ageing 1 . After initial lesion recognition by XPC in global genome repair or by stalled RNA polymerases in transcription-coupled repair, a lesion and surrounding DNA duplex are unwound by TFIIH, which includes the ATPases XPB and XPD, and additional NER factors XPA, XPF, XPG and RPA, to form a DNA bubble 2 comprising around 27 nucleotides. The double strand-single strand (ds-ss) junction-specific endonucleases XPF and XPG cleave DNA on the 5' and 3' sides of the lesion, respectively. Here we report the functional steps and atomic structures of the ATPase-driven and lesion-dependent DNA bubble formation and arrangement of the complete NER factors for dual incision. The unwinding of nearly 30 base pairs of DNA depends mainly on the double strand DNA translocase XPB and the duplex dividers XPA and XPF. XPD binds the lesion strand with XPF at the 5' ds-ss junction. XPF cuts the lesion strand only after XPG binds the 3' ds-ss junction. The ERCC1 subunit of XPF facilitates DNA strand separation and recruitment of RPA to the non-lesion strand. These findings provide insights on the causes of human diseases and potential targets for enhancing chemotherapeutic efficacy.
- Laboratory of Molecular Biology, NIDDK, National Institutes of Health, Bethesda, MD, USA.
Organizational Affiliation: