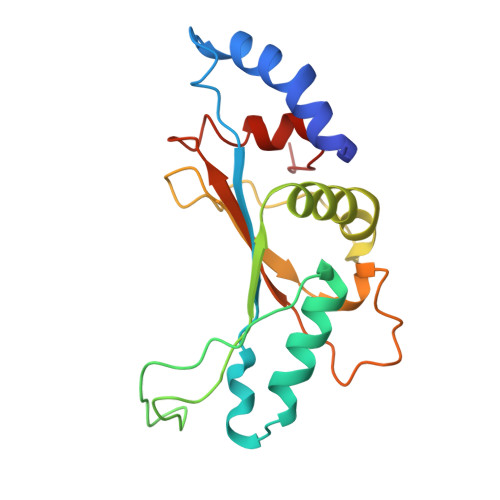
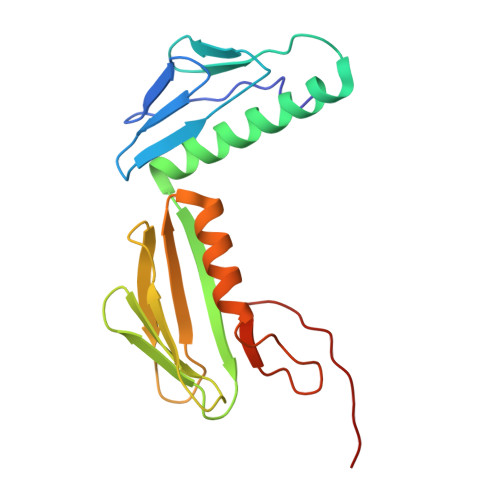
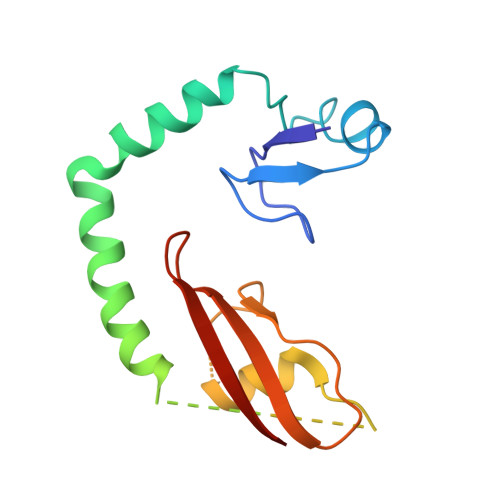
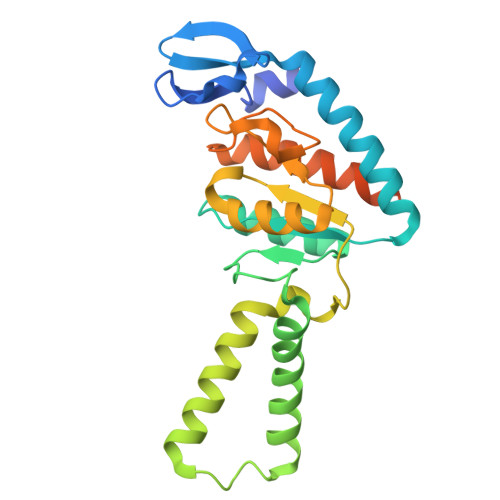
Structure of a specialized 70S initiation complex.
Arpin, D., Roy, B., Naeem, F.M., Fredrick, K., Ortega, J.(2026) Nucleic Acids Res 54
- PubMed: 41854077 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkag094
- Primary Citation Related Structures:
9OY8, 9OYV, 9OZ9, 9OZN, 9OZZ, 9P0E, 9ZCD - PubMed Abstract:
Bacteria of class Bacteroidia lack Shine-Dalgarno (SD) sequences and instead rely on other messenger RNA (mRNA) features, including upstream adenines, for start codon selection. Bacteroidia ribosomes contain the anti-SD (ASD) sequence of 16S ribosomal RNA (rRNA) but are "blind" to SD sequences. This occurs due to the sequestration of the ASD through interactions with bS21, bS18, and bS6 on the 30S platform domain. In many Bacteroidia, including Flavobacterium johnsoniae, there is one gene with an extended SD-rpsU, which encodes bS21. Ribosomes lacking bS21 exhibit high-level translation of rpsU, establishing an autoregulatory circuit in the cell. In this work, we investigate the structural basis of initiation on rpsU mRNA. We find using cryo-electron microscopy that initiation entails the formation of a 13-base pair SD-ASD helix that sterically occludes bS21. Mutations of bS21, bS18, or bS6 that compromise the platform pocket liberate the 3' tail of 16S rRNA, enable SD-ASD pairing, and enhance initiation. As initiation on rpsU mRNA depends on SD-ASD pairing, we infer that dissociation of bS21 from replete ribosomes limits their initiation rate. This work shows how a compositional change of the ribosome can govern translation of a specific gene.
- Department of Anatomy and Cell Biology, McGill University, Montreal, Quebec H3A 0C7, Canada.
Organizational Affiliation: