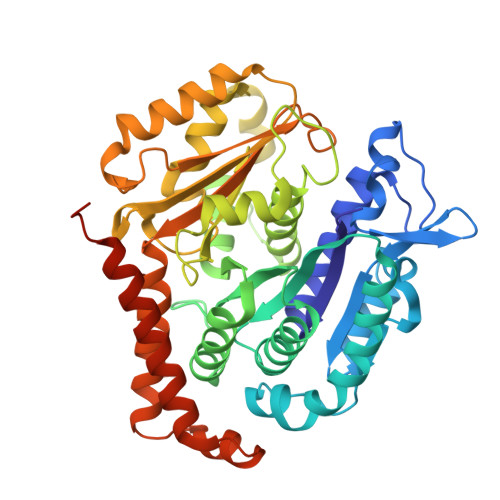
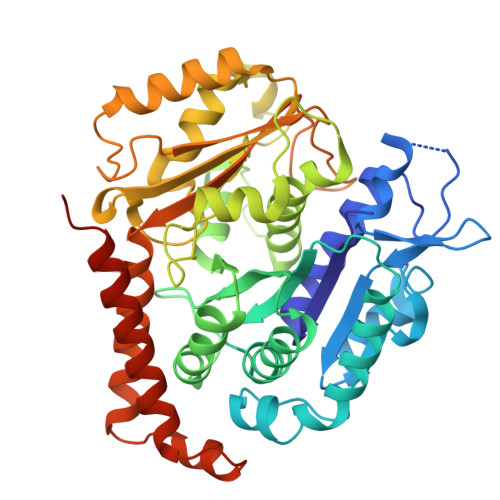
Microtubules in the axon are GDP bound but adopt a stable GTP-like expanded state.
Zehr, E.A., Sun, S., Sarbanes, S.L., Roll-Mecak, A.(2026) Nat Struct Mol Biol 33: 631-640
- PubMed: 41951886 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-026-01787-7
- Primary Citation Related Structures:
9OX7 - PubMed Abstract:
Microtubules scaffold cells, supporting signaling and cargo transport. They assemble from GTP-tubulin, which hydrolyzes to GDP-tubulin during polymerization. GTP-microtubule lattices are stable; GDP lattices depolymerize rapidly. In vitro, hydrolysis triggers lattice compaction. Lattice spacing regulates motors and microtubule-associated proteins; however, the conformation of tubulin in microtubules in cells is unknown. Here, we present the atomic-resolution cryo-electron microscopy structure of human microtubules in situ, in the axons of human cortical neurons derived from induced pluripotent stem cells (iPS cells). Our 2.7-Å-resolution reconstruction delineates bound water molecules and reveals that axonal microtubules adopt an expanded GTP-like lattice, despite being GDP bound. Using cryo-electron tomography and power spectrum analysis, we find that, unlike in axons, microtubules in undifferentiated iPS cells are compacted. Therefore, lattice expansion is part of neuronal differentiation. Our work provides molecular insights into neurogenesis and has implications for understanding microtubule stability and effector recruitment in neurons.
- Cell Biology and Biophysics Unit, National Institute of Neurological Disorders and Stroke, Bethesda, MD, USA.
Organizational Affiliation: