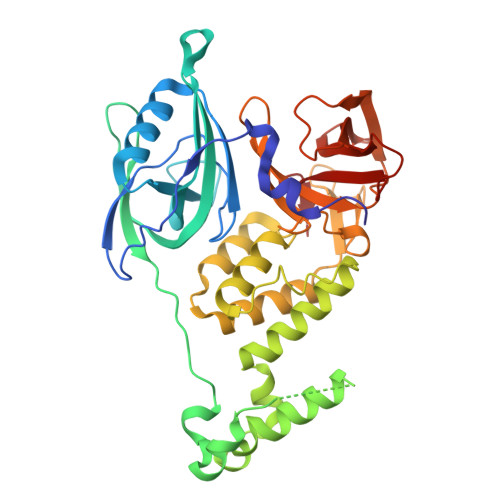
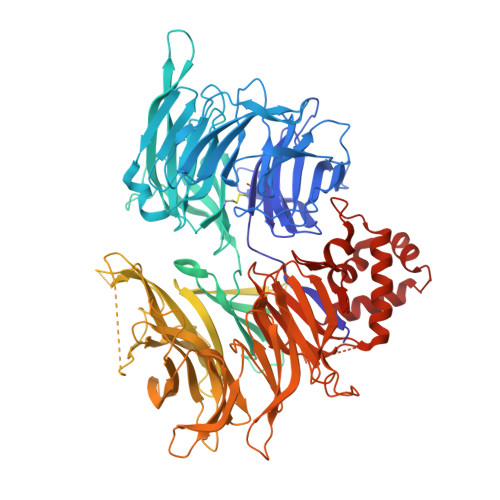
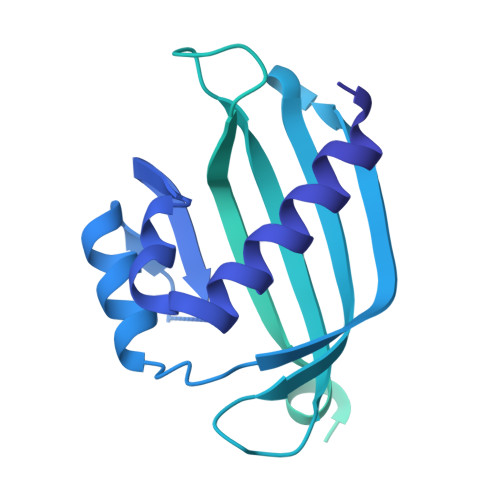
Cereblon induces G3BP2 neosubstrate degradation using molecular surface mimicry.
Annunziato, S., Quan, C., Donckele, E.J., Lamberto, I., Bunker, R.D., Zlotosch, M., Schwander, L., Murthy, A., Wiedmer, L., Staehly, C., Matysik, M., Gilberto, S., Kapsitidou, D., Wible, D., De Donatis, G.M., Trenh, P., SriRamaratnam, R., Strande, V., Lieberherr, R., Lyon, D., Steiner, D., Silva, J., Almeida, R., Dolgikh, E., DeMarco, B., Tsai, J., Sadok, A., Zarayskiy, V., Walter, M., Tiedt, R., Lumb, K.J., Bonenfant, D., Fasching, B., Castle, J.C., Townson, S.A., Gainza, P., Petzold, G.(2026) Nat Struct Mol Biol 33: 479-487
- PubMed: 41559416 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-025-01738-8
- Primary Citation Related Structures:
9OS2 - PubMed Abstract:
Molecular glue degraders (MGDs) are small-molecule compounds that divert E3 ligases to degrade nonnatural substrates called neosubstrates. Clinically effective MGDs bind cereblon (CRBN), a substrate receptor of the Cullin 4-RING E3 ubiquitin ligase (CRL4 CRBN ), and recruit neosubstrates to an MGD-induced neosurface on the CRBN CULT domain through molecular mimicry of a natural CRBN degron. Here, we identify G3BP2 (Ras-GAP SH3 domain-binding protein 2), a neosubstrate that bypasses canonical interactions with CRBN by engaging an unconventional binding site on the CRBN LON domain. The ternary complex interface does not resemble known interactions with CRBN. Instead, CRBN leverages a preexisting protein-protein interaction (PPI) hotspot on the target protein by mimicking an endogenous binding partner of G3BP2. Our findings suggest that composite neosurfaces that mimic and stabilize the footprint of natural PPIs (in short, 'glueprints') could become a viable strategy for the rational expansion of the MGD target repertoire.
- Monte Rosa Therapeutics, Inc., Boston, MA, USA.
Organizational Affiliation: