Rapidly acquired HIV-1 neutralization breadth in a rhesus V2 apex knockin mouse model after a single bolus immunization.
Ghosh, A.R., Habib, R., Mishra, N., Roark, R.S., Akauliya, M., Albowaidey, A.A., Allen, J.D., Amereh, K., Avillion, G., Bottermann, M., Liang, B., Chaudhary, N., Callaghan, S., Dye, J., Li, X., Ellis-Pugh, J.R., Roy Chowdhury, R., James, N.E., Liu, X., Maiorino, L., Villavicencio, P.M., Nedellec, R., Oberoi, P., Sowers, K.J., Park, Y., Prum, T., Rodriguez, L., Ssozi, M., Torres, J.L., Walsh, A.A., Warner, J.E., Weldon, S.R., Xu, L., Wiehe, K., Crispin, M., Ward, A.B., Nair, U., Hahn, B.H., Burton, D.R., Shapiro, L., Kwong, P.D., Irvine, D.J., Andrabi, R., Shaw, G.M., Batista, F.D.(2026) Sci Immunol 11: eadz5064-eadz5064
- PubMed: 41686913 Search on PubMed
- DOI: https://doi.org/10.1126/sciimmunol.adz5064
- Primary Citation Related Structures:
9OOG, 9OOK, 9OOM - PubMed Abstract:
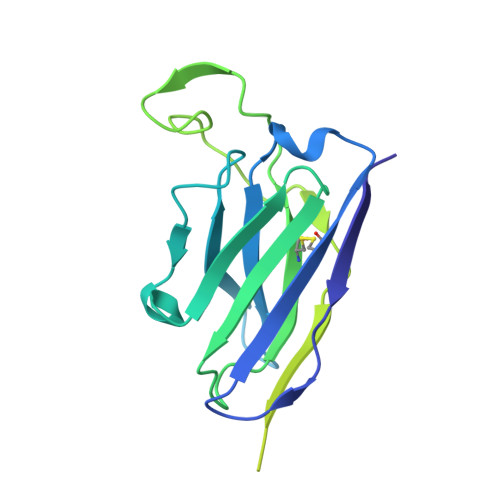
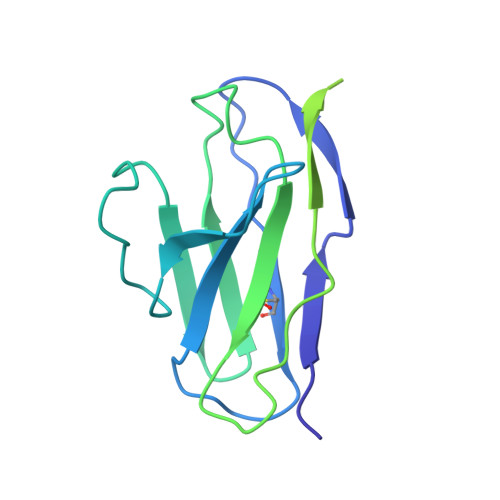
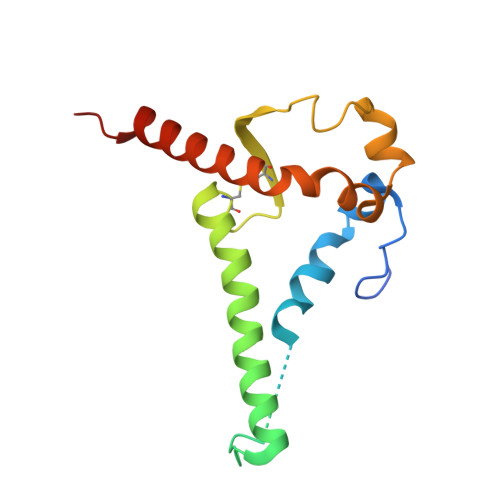
Current immunization strategies to elicit broadly neutralizing antibodies (bnAbs) against HIV-1 generally propose complex, multiboost regimens. In rhesus macaques, simian-human immunodeficiency virus (SHIV) infection rapidly drives the development of some bnAb classes sharing structural similarities with those in humans. Here, we generated a knockin (KI) mouse model with B cells bearing the unmutated common ancestor of a V2 apex-targeted bnAb lineage, V033-a. A single immunization with a germline-targeting native-like trimer, Q23-APEX-GT1, recapitulated the ontogeny of the mature rhesus bnAb in KI mice, including rare, disfavored somatic mutations. Resulting antibodies exhibited potent neutralization against a broad panel of heterologous HIV-1 strains. Boosting with Env escape mutant trimers further improved breadth and potency, and cryo-electron microscopy analysis revealed the structural basis for heterologous neutralization breadth. Nonhuman primate and mouse models combined with structure can serve as a platform for identifying and validating immunogens that streamline HIV vaccination regimens.
- Batista Lab, Ragon Institute of Mass General Brigham, MIT, and Harvard, Cambridge, MA 02139, USA.
Organizational Affiliation: