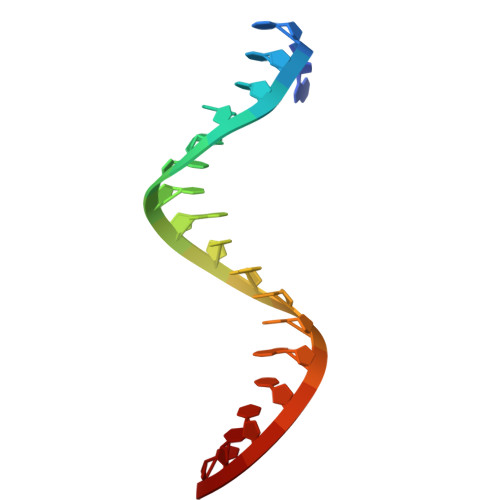
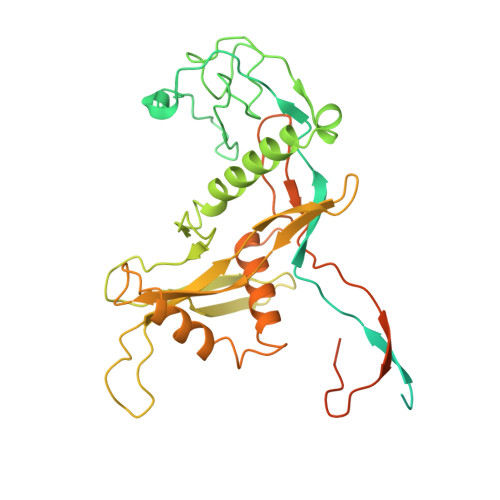
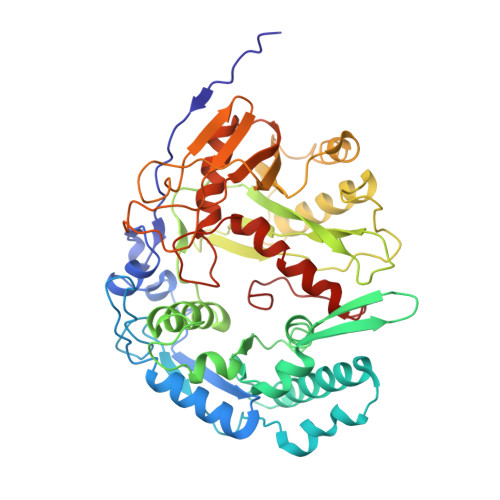
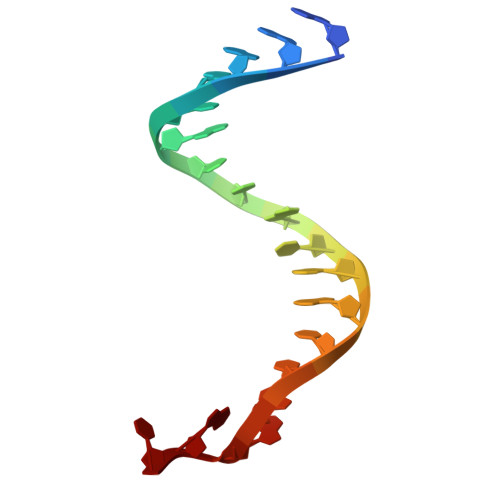
Argonaute-HNH filaments triggered by invader DNA confer bacterial immunity.
Kanevskaya, A., Narwal, M., Lisitskaya, L., Moiseenko, A.V., Sokolova, O.S., Sluchanko, N.N., Murakami, K.S., Kulbachinskiy, A.(2025) Nat Commun 16: 11389-11389
- PubMed: 41315870 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-66189-7
- Primary Citation Related Structures:
9OM4 - PubMed Abstract:
Argonaute proteins provide innate immunity in all domains of life through guide-dependent recognition and cleavage of invader nucleic acids. Many short prokaryotic Argonautes (pAgos) lack nuclease activity and are instead co-encoded with tentative nuclease effectors, but their activation mechanisms remain unknown. Here, we characterize SPARHA systems (short prokaryotic argonautes, HNH-associated), containing HNH nuclease effectors. RNA-guided target DNA recognition by pAgo induces formation of SPARHA filaments with a chain of double nuclease sites formed by HNH tetramers, leading to indiscriminate collateral degradation of DNA and protecting bacterial population from invaders. We show that the assembly of filaments proceeds via a universal activation pathway involving a cascade of target-induced conformational changes in SPARHA, conserved in other short pAgo systems containing various types of effectors. pAgos and associated effectors act as modular defense systems that translate recognition of specific DNA into immune response through assembly of supramolecular complexes, deleterious for invaders and potentially useful for biotechnology.
- Institute of Gene Biology, Russian Academy of Sciences, Moscow, Russia. annakanevskaya5@gmail.com.
Organizational Affiliation: