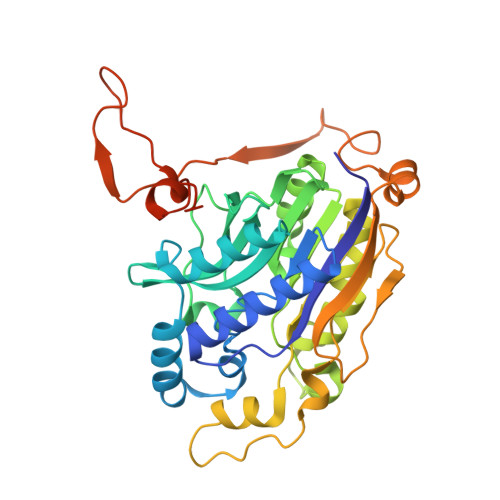
The single-particle cryo-EM structures of a bacterial cyanide dihydratase and a fungal cyanide hydratase.
Justo Arevalo, S., Valle-Riestra Felice, V., Barahona Acuna, M., Ordinola Flores, K., Quinones Aguilar, M., Balan, A., Farah, C.S.(2026) Structure 34: 599-610.e2
- PubMed: 41709456 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2026.01.009
- Primary Citation Related Structures:
9ODT, 9OFA - PubMed Abstract:
Cyanide is widely used in industries due to its affinity for metals, a property that also underlies its toxicity. Industries, therefore, must reduce cyanide concentration before the final disposal of wastewater. Physical, chemical, and biological methods have been developed for this; however, knowledge about the structure of enzymes involved in cyanide degradation remains limited. Structural characterization of these proteins could facilitate the development of enzymes with enhanced bioremediation potential. Here, we present the single-particle cryo-electron microscopy structures of a cyanide dihydratase from Bacillus safensis and a cyanide hydratase from Gloeocercospora sorghi at 2.2 Å and 2.0 Å resolution, respectively. We provide a comprehensive description and comparative analysis alongside all previously determined nitrilase structures. Importantly, our full-length structures reveal new features in the C-terminal as well as specific intermolecular interactions between protomer interfaces and within the helix lumen. Finally, our findings offer insights into the reaction mechanisms of these two enzymes.
- Departamento de Bioquimica, Instituto de Quimica - Universidade de São Paulo, São Paulo, Brazil; Facultad de Ciencias Biologicas - Universidad Ricardo Palma, Santiago de Surco, Peru. Electronic address: santiago.jus.are@usp.br.
Organizational Affiliation: