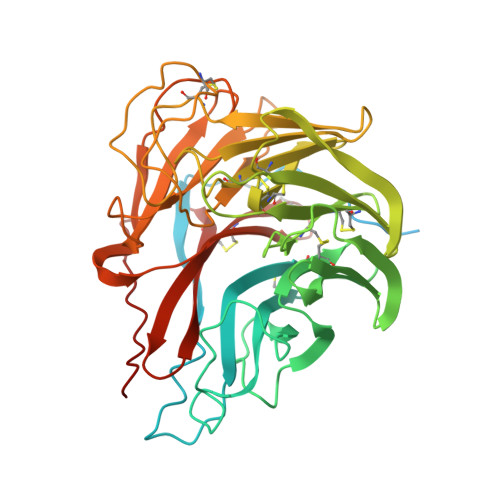
Characterization of the glycoproteins of novel fish influenza B-like viruses.
Singh, G., Huang, J., Bhavsar, D., Vasilev, K., Ferguson, J.A., Boons, G.J., Simon, V., de Vries, R.P., Han, J., Ward, A., Krammer, F.(2025) bioRxiv
- PubMed: 40654808 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1101/2025.05.08.652883
- Primary Citation Related Structures:
9O5U, 9O5W - PubMed Abstract:
Novel influenza-like virus sequences previously identified in fish and amphibians were found to cluster as a sister clade of influenza B viruses, but have thus far remained uncharacterized. We demonstrate that salamander influenza-like virus (SILV) HA is functionally divergent from influenza B virus HA and does not bind to α 2,3- and α2,6-linked sialic acids. However, the HAs of Siamese algae-eater influenza-like virus (SAEILV) and chum salmon influenza-like virus (CSILV) bind to α2,3 linked sialic acid. Furthermore, SAEILV HA binds to sialyated Lewis X, is activated by human airway enzymes and is fusogenic at a wide range of pH conditions. SAEILV NA has a highly conserved active site and a similar structure to other known NAs. We also determined the cryo-electron microscopy structure of the HA of a previously described virus from the same sister clade, the Wuhan spiny eel influenza virus (WSEIV). Importantly, no cross-reactive antibodies against these HAs or NAs were found in the human serum, suggesting that humans are immunologically naïve to these viruses.
- Department of Microbiology, Icahn School of Medicine at Mount Sinai, New York, NY, USA.
Organizational Affiliation: