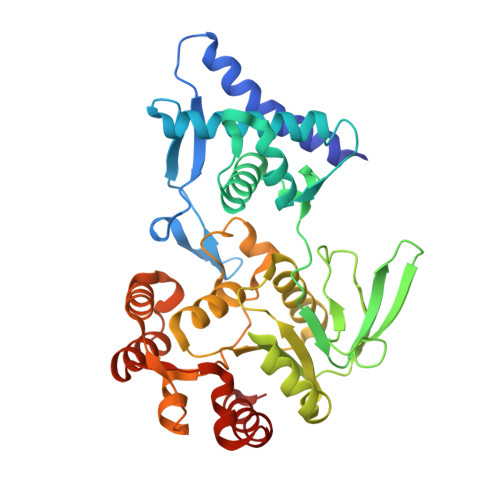
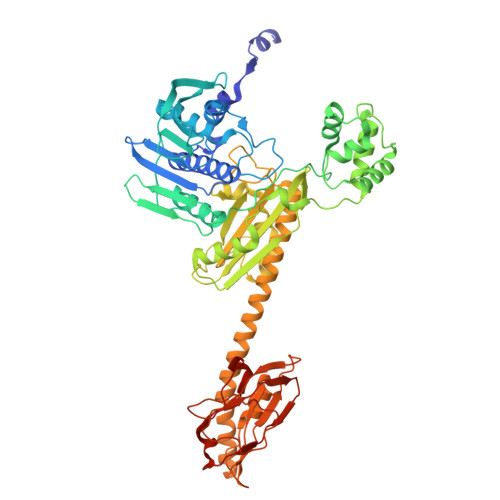
Supercoiled DNA recognition and cleavage control in topoisomerase VI.
Richman, D.E., Wendorff, T.J., Rashid, F., Beck, C., Yan, Q., Johnson, H.R., Eckerty, R.A., Fogg, J.M., Baker, M.L., Zechiedrich, L., Berger, J.M.(2026) Nat Commun 17
- PubMed: 41698922 Search on PubMed
- DOI: https://doi.org/10.1038/s41467-026-69491-0
- Primary Citation Related Structures:
9O0G, 9O7O, 9O8P, 9O8Z, 9O9M - PubMed Abstract:
Type II topoisomerases modulate DNA supercoiling and resolve chromosome entanglements. Type IIB topoisomerases, exemplified by DNA topoisomerase VI (Top6), are used by plants and archaea to support endoreduplication and cell proliferation, respectively; homologs of Top6 further serve to initiate meiotic recombination in eukaryotes and constitute the nuclease portion of MksBEFG/Wadjet/Gabija bacterial defense systems. To understand how such factors act upon DNA, we determine structures of Top6 bound to supercoiled minicircles in cleaved and uncleaved states using single-particle electron cryo-microscopy. The structures show that Top6 binds a curved 74 bp region of the supercoiled minicircle DNA and that it cuts at a distinct deformability motif, explaining its preference for supercoiled substrates and highlighting the role of DNA plasticity in cleavage site selection. Dynamic protein-DNA interactions and an unanticipated tension sensor help recognize bent DNA and couple ATPase disposition to cleavage state activation. Our observations explain how DNA recognition and cleavage by type II topoisomerases are regulated by interdependent structural changes in DNA and the enzyme.
- Department of Biophysics & Biophysical Chemistry, Johns Hopkins University School of Medicine, Baltimore, MD, USA.
Organizational Affiliation: