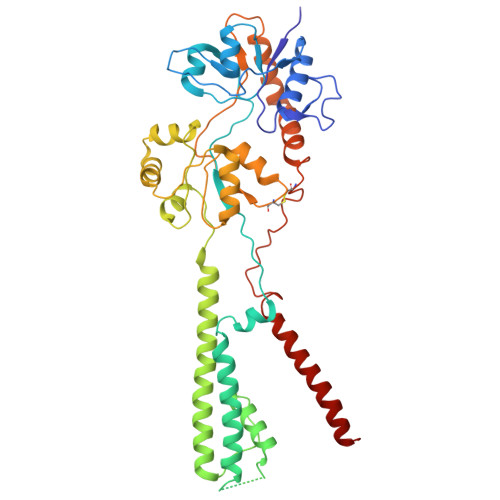
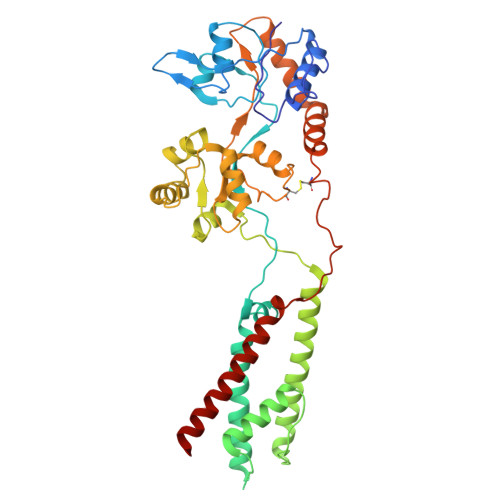
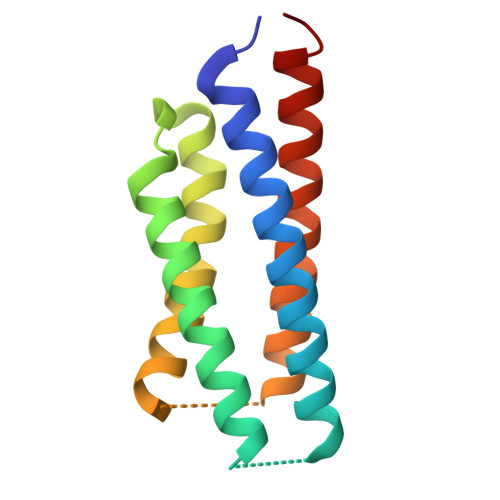
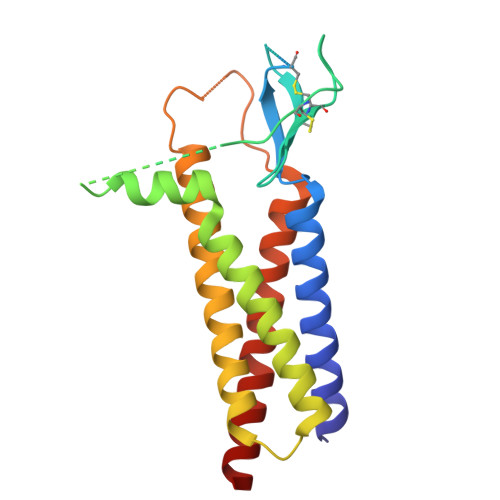
Gating and noelin clustering of native Ca 2+ -permeable AMPA receptors.
Fang, C.L., Spangler, C.J., Park, J., Sheldon, N., Trussell, L.O., Gouaux, E.(2025) Nature 645: 526-534
- PubMed: 40550474 Search on PubMed
- DOI: https://doi.org/10.1038/s41586-025-09289-0
- Primary Citation Related Structures:
9NR6, 9NR7, 9NR8, 9NR9, 9NRA - PubMed Abstract:
AMPA-type ionotropic glutamate receptors (AMPARs) are integral to fast excitatory synaptic transmission and play vital roles in synaptic plasticity, motor coordination, learning, and memory 1 . While extensive structural studies have been conducted on recombinant AMPARs and native calcium impermeable (CI)-AMPARs alongside their auxiliary proteins 2-5 , the molecular architecture of native calcium permeable (CP)-AMPARs has remained undefined. To elucidate the subunit composition, physiological architecture, and gating mechanisms of CP-AMPARs, here we present the first visualization of these receptors, immunoaffinity purified from rat cerebella, and resolve their structures using cryo-electron microscopy (cryo-EM). Our results indicate that the predominant assembly consists of GluA1 and GluA4 subunits, with the GluA4 subunit occupying the B and D positions, while auxiliary subunits, including TARPs, are located at the B'/D' positions and CNIHs or TARPs at the A'/C' positions. Furthermore, we resolved the structure of the Noelin 1-GluA1/A4 complex, wherein Noelin 1 (Noe 1) specifically binds to the GluA4 subunit at the B and D positions. Notably, Noe 1 stabilizes the amino-terminal domain (ATD) layer without affecting receptor gating properties. Noe 1 contributes to AMPAR function by forming dimeric-AMPAR assemblies that likely engage in extracellular networks, clustering receptors within synaptic environments and modulating receptor responsiveness to synaptic inputs.
- Vollum Institute, Oregon Health & Science University, Portland, OR, USA.
Organizational Affiliation: