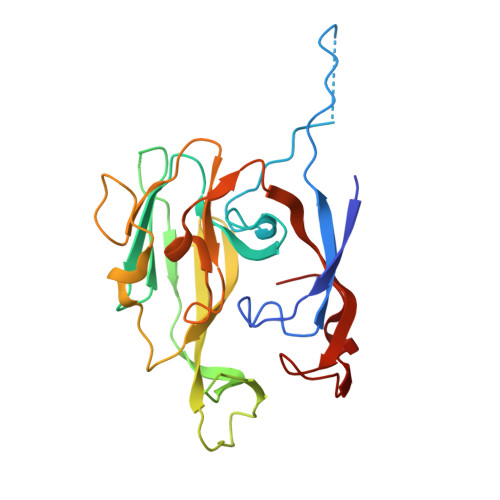
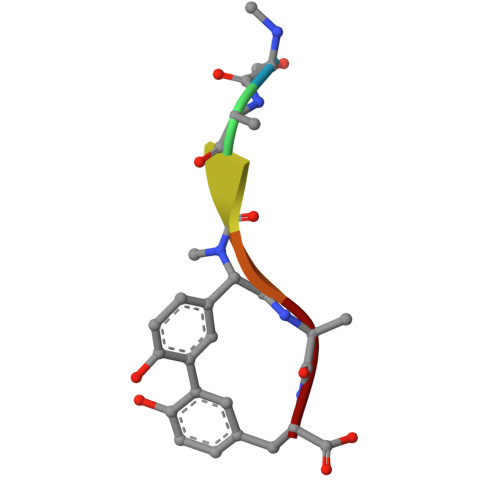
Crystal structure of Escherichia coli type I signal peptidase P84A in complex with lipopeptide antibiotic arylomycin A 2 .
Luo, C., Paetzel, M.(2025) J Struct Biol 217: 108260-108260
- PubMed: 41207487 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2025.108260
- Primary Citation Related Structures:
9NLO - PubMed Abstract:
Type I signal peptidase (SPase I) is an essential membrane-bound enzyme that removes amino-terminal signal peptides from secretory proteins. Owing to its critical role in bacterial viability and its periplasmic accessibility, SPase I has emerged as an attractive target for antibiotic development. Arylomycins, a class of macrocyclic lipohexapeptide natural products, inhibit SPase I by binding to its active site. Previous studies have identified a key resistance determinant-a proline residue at the base of the substrate-binding groove (Pro84 inEscherichia coliSPase I)-which reduces arylomycin affinity. Here, we present the crystal structure of theE. coliSPase I P84A mutant in complex with arylomycin A 2 , revealing that the introduced alanine enables an additional hydrogen bond between the enzyme backbone and the arylomycin N-terminal carbonyl, thus enhancing the affinity for arylomycins. Furthermore, a newly developed preprotein-binding assay utilizing a non-cleavable version of ProOmpA Nuclease A demonstrates that substituting SPase I Pro84 with serine or leucine disrupts substrate recognition, underscoring the delicate balance between inhibitor resistance and substrate processing. These findings reveal that residue Pro84 participates in the interaction between preprotein signal peptides and the E. coli SPase I substrate-binding groove, offering a foundation for designing next-generation arylomycin analogs with improved antibacterial potency.
- Department of Molecular Biology and Biochemistry, Simon Fraser University, South Science Building 8888 University Drive, Burnaby, British Columbia V5A 1S6 Canada.
Organizational Affiliation: