Engineered antibodies that stabilize drug-modified KRAS G12C neoantigens enable selective and potent cross-HLA immunotherapy.
Maso, L., Mosure, S.A., Rodriguez-Aponte, S.A., Pizzo, A., Mensah, D.N., Southard, M., Sze, S., Ahmed, T., Vash, B., Hattori, T., Rajak, E., Koide, A., Neel, B.G., Koide, S., Liu, W., Toenjes, S.T., Jardine, P.D.S., Chopra, R., Rader, C., Stopfer, L.E.(2025) Nat Commun 16: 11264-11264
- PubMed: 41408054 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-66132-w
- Primary Citation Related Structures:
9NFB, 9NFC - PubMed Abstract:
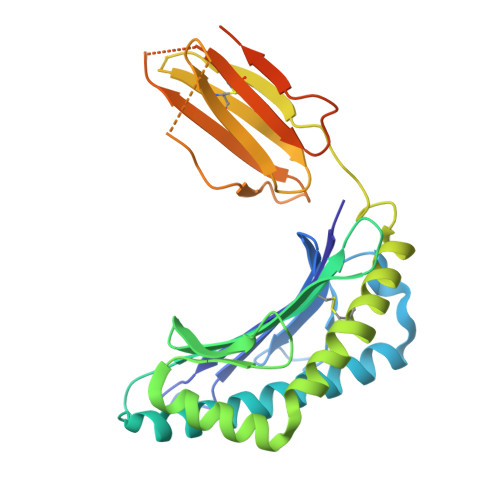
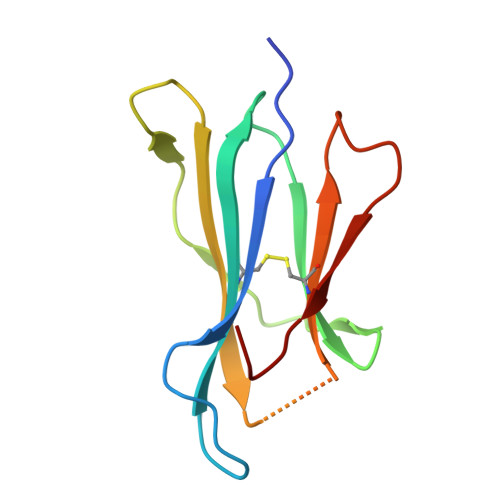
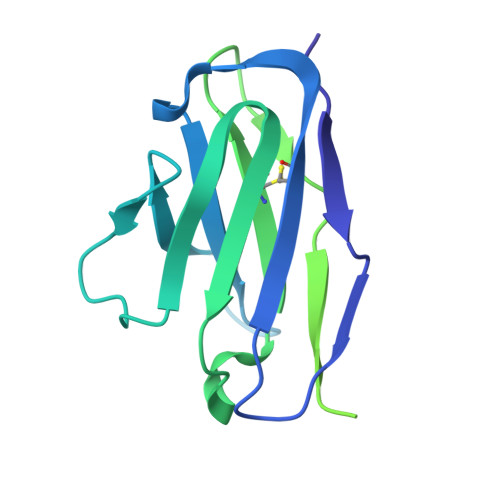
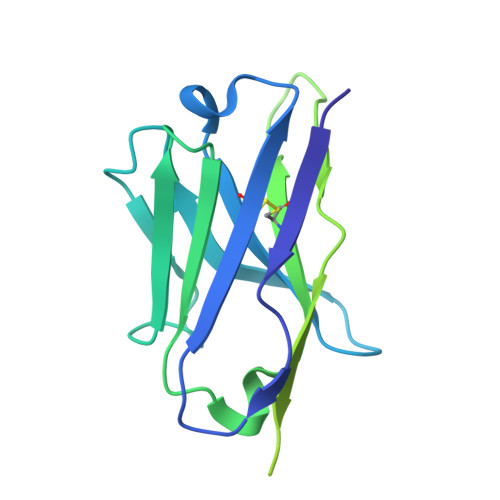
Covalent inhibitors of oncoprotein KRAS have initial efficacy, but responses lack durability. Covalently modified oncoproteins are presented as MHC-restricted hapten-peptides (p*MHC) on the cancer cell surface, enabling combination of targeted therapy with immunotherapy to overcome drug resistance. Building on indirect evidence of KRAS G12C -derived p*MHCs, we use immunopeptidomics to identify and directly quantify these synthetic neoantigens. To address challenges by their low copy number, we develop AETX-R114, a T cell engaging bispecific antibody with picomolar affinity for MHC-restricted sotorasib-modified KRAS G12C peptides presented by three HLA-A3 supertype alleles. AETX-R114 dramatically increases the half-life and thereby the number of presented p*MHCs, enabling selective and potent killing of resistant cancer cells both in vitro and in vivo. To broaden the therapeutic potential of creating and targeting synthetic neoantigens, we further develop AETX-R302, which recognizes divarasib-modified KRAS G12C peptides presented on alleles from the HLA-A2 and A3 supertypes. Cryo-EM structure determination reveals the molecular basis for breaking HLA supertype restriction. Collectively, our study illustrates how engineered antibodies can transform synthetic neoantigens into actionable cancer immunotherapy targets.
- Aethon Therapeutics, Long Island City, NY, USA.
Organizational Affiliation: