Structural and mechanistic insights into herpesvirus helicase-primase and its therapeutic inhibitors.
Yao, Q., Mercier, A., Nayak, A., May, L., Ho, P.Y., Lewis-Ballester, A., Nair, V., Sapre, A., Aeschbacher, T., Mukherjee, J., Richards, C., Mateo, R., Cho, A., Lansdon, E., Yu, X.(2025) Nat Microbiol 10: 3191-3201
- PubMed: 41188384 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41564-025-02168-4
- Primary Citation Related Structures:
9NDA, 9NDQ, 9NDT, 9NDZ, 9NE0, 9NEB, 9NEE, 9NEL - PubMed Abstract:
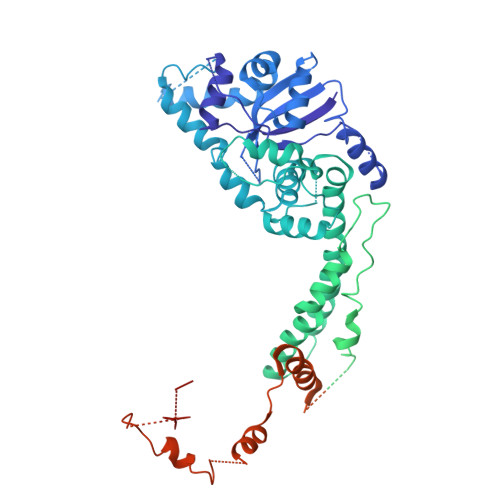
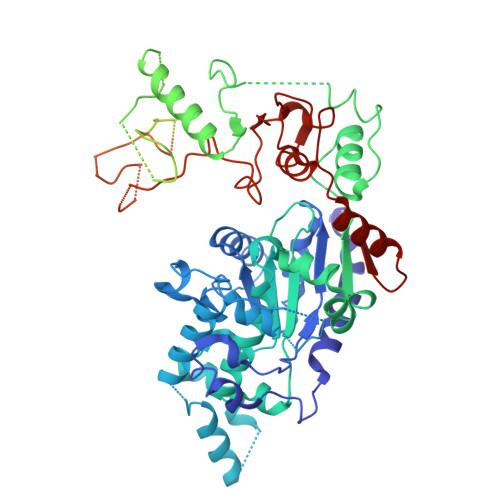
The herpes simplex virus (HSV) helicase-primase (HP) complex is a promising anti-herpes therapeutic target. However, progress in developing highly effective small-molecule HP inhibitors (HPIs) for the treatment of genital herpes has been hindered by the lack of structural information on the HP complex and the incomplete understanding of the mechanism of action of HPIs. Here we present the cryogenic electron microscopy structure of the HSV-1 HP apo-complex (3.8 Å), along with structures bound to pritelivir (3.2 Å) and amenamevir (3.2 Å)-two clinically active, chemically distinct HPIs. The potency of both inhibitors against HSV variants bearing mutations within the HPI binding pocket supports the high-resolution mapping of key molecular interactions while revealing residues that govern their antiviral spectrum against alphaherpesviruses. Our results provide important insight into the unique architecture of the HP complex and the mechanism of inhibition of HPIs, paving the way for the development of next-generation antivirals to treat herpesvirus infections.
- Gilead Sciences Inc., Foster City, CA, USA. qing.yao1@gilead.com.
Organizational Affiliation: