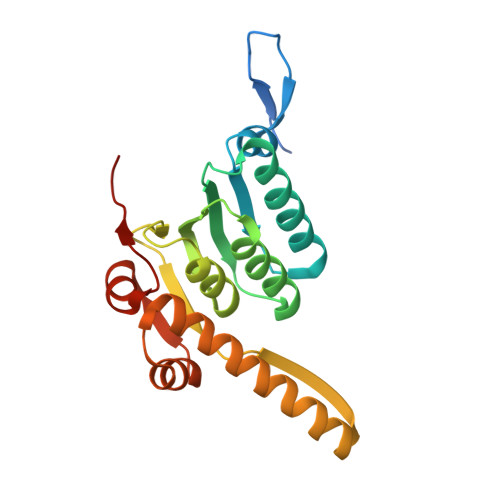
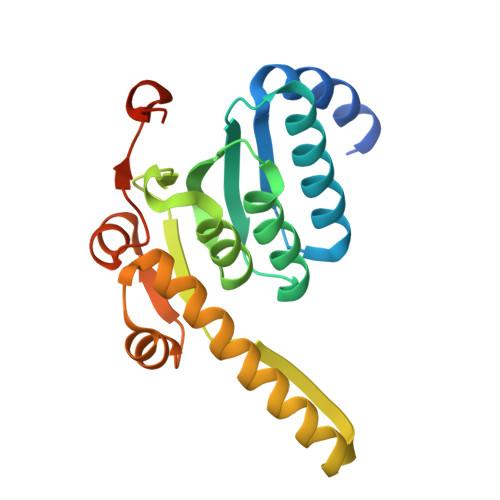
Structural insights into the Pseudomonas aeruginosa ClpP1•ClpP2 heterocomplex and its interactions with the AAA+ ClpX unfoldase.
Ghanbarpour, A., Zhang, J.J., Davis, J.H., Baker, T.A., Sauer, R.T.(2025) Protein Sci 34: e70310-e70310
- PubMed: 40980994 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.70310
- Primary Citation Related Structures:
9NDJ - PubMed Abstract:
ClpXP and other AAA+ proteases play central roles in bacterial proteostasis by degrading misfolded and regulatory proteins. In Pseudomonas aeruginosa, ClpXP consists of the ClpX unfoldase and ClpP peptidase, which influence critical adaptive processes contributing to stress resistance. P. aeruginosa Pa ClpP1 and Pa ClpP2 paralogs assemble into homomeric ( Pa ClpP1•ClpP1) and heteromeric ( Pa ClpP1•ClpP2) complexes. Pa ClpP2 is only active in the Pa ClpP1•ClpP2 heterocomplex. Here, we present a cryo-EM structure of Pa ClpX•ClpP1•ClpP2, revealing how Pa ClpX binds Pa ClpP1, which in turn interacts with Pa ClpP2. Comparison of the active heterocomplex with an inactive Pa ClpP2 crystal structure shows that Pa ClpP1 binding induces conformational changes in Pa ClpP2, stabilizing an active catalytic triad. Differences in Pa ClpP1 and Pa ClpP2 substrate-binding residues and an unstructured ClpP2 N-terminal segment that protrudes into the peptidase chamber likely contribute to distinct peptide-cleavage specificities of Pa ClpX•ClpP1•ClpP2 and Pa ClpX•ClpP1•ClpP1. Given the role of Pa ClpP1•ClpP2 in biofilm formation and virulence, these structural insights may provide a foundation for developing selective inhibitors to combat P. aeruginosa infections.
- Department of Biochemistry and Molecular Biophysics, Washington University in St. Louis, St. Louis, Missouri, USA.
Organizational Affiliation: