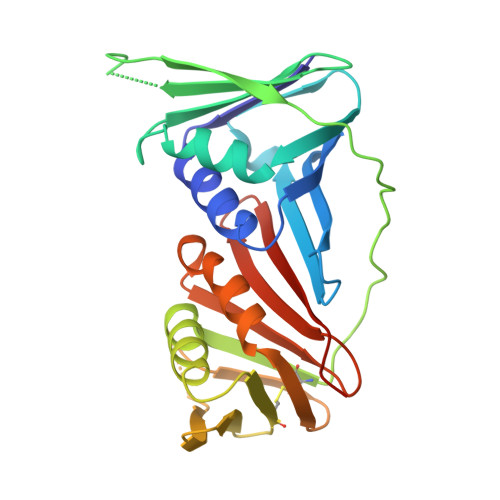
Screening for novel chemical scaffolds targeting PCNA identifies the Hsp90alpha inhibitor SNX-2112.
Jossart, J., Ma, N., Haratipour, P., Li, C.M., Jang, C., Gu, L., O'Brien, T.E., Vaidehi, N., Malkas, L.H., Hickey, R.J., Perry, J.J.P.(2026) Curr Res Struct Biol 11: 100183-100183
- PubMed: 41743503 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.crstbi.2026.100183
- Primary Citation Related Structures:
9N3L - PubMed Abstract:
Proliferating cell nuclear antigen (PCNA) protein is an emerging therapeutic target for several diseases including cancers, and certain viral infections. It was previously regarded as being undruggable, due to its functions as a molecular hub that regulates a multitude of essential cellular pathways. However, we recently developed a PCNA inhibitor, AOH1996, which acts as a molecular glue between PCNA and RNA Pol II promoting selective killing of cancer cells through the induction of transcription-replication conflicts. We have now moved this investigational new drug into Phase 1 clinical trials. Yet, the discovery of new PCNA inhibitors is still of notable importance, because of the potential for further improving the potency and overall in vivo stability relative to AOH1996, and to potentially provide more treatment options for differing patient populations and for overcoming drug resistance mechanisms that may arise from AOH1996 treatment. Moreover, targeting other PCNA mediated disease states could require the identification of novel compounds that have non-identical mechanisms of action or differing pharmokinetic and pharmodynamic profiles. We coupled artificial intelligence-based computational screens with thermal shift assays in a search for novel hit compounds, and characterized a key hit through protein crystallography, a cellular thermal shift assay and protein frustration calculations. Notably, we discovered that the Hsp90α inhibitor, SNX-2112, binds to PCNA within the major PIP-box binding pocket, revealing a new chemical scaffold for the potential development of novel PCNA-targeting inhibitors.
- Department of Cancer Biology and Molecular Medicine, City of Hope Medical Center, Monrovia, CA, USA.
Organizational Affiliation: