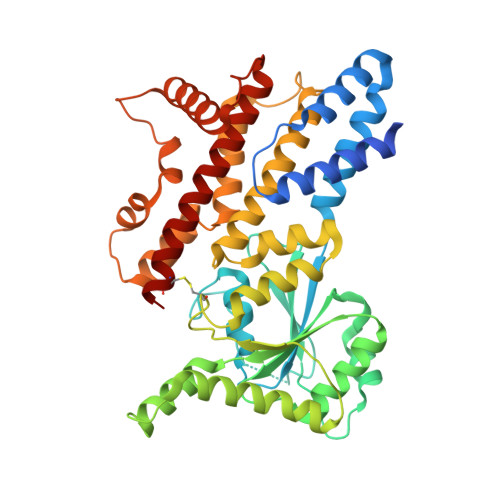
Structural insights into GM4951 as a lipid droplet GTPase regulating hepatic lipid metabolism.
Raj, R., Jiang, Y., Jha, R.K., Moresco, E.M.Y., Joshi, H., Zhang, Z., Beutler, B.(2025) Nat Commun 16: 11458-11458
- PubMed: 41387427 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-025-66253-2
- Primary Citation Related Structures:
9N2Q, 9N2R, 9N2S, 9N2T, 9N6D - PubMed Abstract:
GM4951 is an immunity-related GTPase (IRG) that counteracts hepatic lipid accumulation in mice fed a high-fat diet. We determine full-length protein structures of GTPγS- and GDP-bound GM4951, and two missense mutants (N86K or D125G) associated with metabolic dysfunction-associated steatotic liver disease (MASLD) in mice. All four structures reveal a conserved GTPase domain fold and a helix bundle composed of the N- and C-terminal regions. Each mutation alters the dynamics of the switch-I and switch-II loops important for catalytic function and lipid droplet (LD) localization. GM4951 predominantly forms dimers in vitro. Cryo-electron microscopy reveals a dimer interface formed by the helical domains of two protomers (tail to tail), distinct from other IRGs. The N-terminal helices are necessary for LD localization, while a disulfide bond between helices in the GTPase domain and C-terminus is necessary for interaction with MASLD-associated HSD17B13. Distinct N- and C-terminal conformations set GM4951 apart from other IRGs structurally and functionally.
- Center for the Genetics of Host Defense, University of Texas Southwestern Medical Center, Dallas, TX, USA. Rishi.Raj@UTSouthwestern.edu.
Organizational Affiliation: