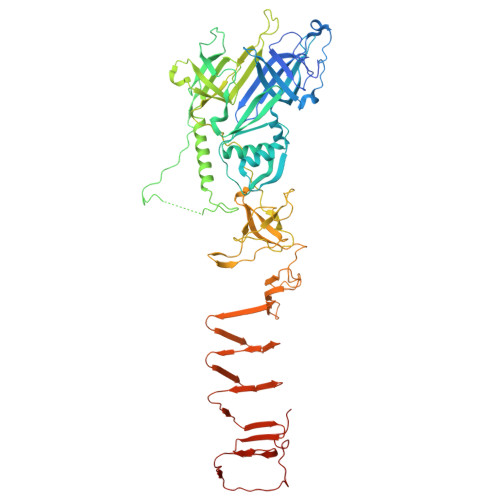
Structure of the type VI secretion protein VgrS from Salmonella Typhimurium.
Sachar, K., Van Schepdael, M., Winters, K.L., Prehna, G.(2026) J Struct Biol 218: 108297-108297
- PubMed: 41687690 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2026.108297
- Primary Citation Related Structures:
9MLP - PubMed Abstract:
Enteric bacterial pathogens employ various strategies to colonize the intestine and cause diseases ranging from gastroenteritis to systemic infections. For example, Salmonella enterica utilizes a nanomachine known as the type VI secretion system (T6SS) to facilitate colonization of the host gut. However, the varied mechanistic details of how the T6SS is loaded with effector proteins remains to be elucidated. Here, we present an X-ray crystal structure of the Salmonella Typhimurium VgrG (VgrS) that serves as platform for T6SS effector loading. Compared to other known structures of VgrG proteins, the VgrS trimer adopts an alternative open conformation within the gp27 region base. The open conformation is due to an extended loop conformation in the gp27 region. This conformation creates a domain extension which docks into the neighboring monomer sequentially around the trimer. Additionally, a comparative structural analysis of VgrS with other VgrG proteins reveals molecular variations that may contribute to specific effector loading mechanisms. Our structural data and molecular analysis highlight the observation that the T6SS of each bacterial species or strain is unique.
- Department of Microbiology, University of Manitoba, Winnipeg, MB R3T 2N2, Canada.
Organizational Affiliation: