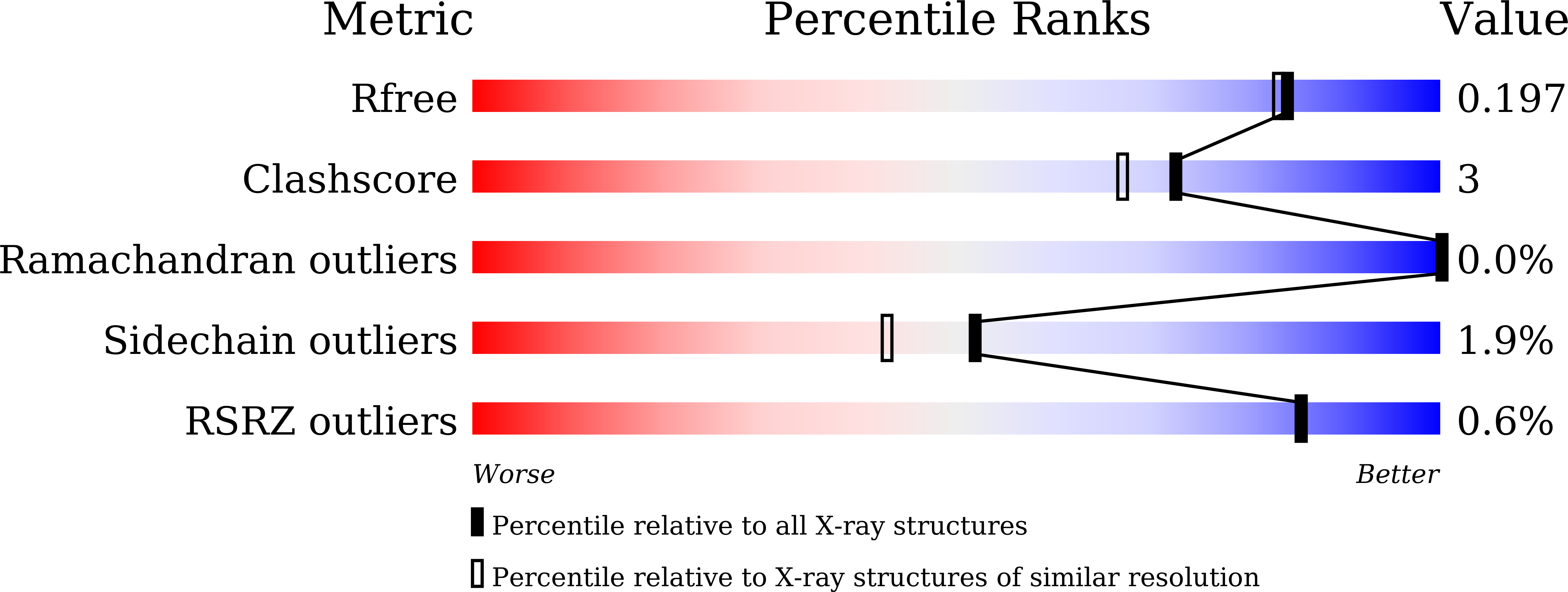
Chemical inhibition of INO1 reduces phytic acid in rice and wheat grains for enhanced micronutrient bioavailability.
Akabane, T., Kamino, S., Okamura, T., Sugaya, A., Nakamura, M., Ikeda, K., Yonezawa, T., Fukushima, A., Kojima, S., Aoki, Y., Chujo, S., Yamauchi, Y., Shibusawa, R., Ishimaru, K., Nagasaka, S., Katoh, E., Hirotsu, N.(2026) Nat Food 7: 163-173
- PubMed: 41639471
- DOI: https://doi.org/10.1038/s43016-026-01295-3
- Primary Citation Related Structures:
9M53 - PubMed Abstract:
Phytic acid (PA), the major phosphorus storage compound in cereal grains, chelates essential minerals such as iron and zinc, thereby preventing their absorption in the human intestine and contributing to micronutrient deficiencies. Myo-inositol 3-phosphate synthase 1 (INO1) is a key enzyme in PA biosynthesis, catalysing the pathway's first step and determining PA contents in cereal grains. Although the genetic modification of PA biosynthesis enzymes can reduce PA levels in grains by disrupting enzyme function, it often impairs germination and seedling growth. Here we used a plant chemical biology approach targeting INO1 to reduce PA levels in rice (Oryza sativa L.) and wheat (Triticum aestivum L.) through chemical intervention. From over 1,000 molecules, candidate compounds were identified on the basis of biophysical or biochemical screening. When applied to developing seeds in panicles or spikes, these selected compounds reduced the PA content in grains of rice and wheat. This study presents a strategy for developing low-PA crops and contributes to global efforts to reduce malnutrition.
- Graduate School of Life Sciences, Toyo University, Asaka, Japan.
Organizational Affiliation: