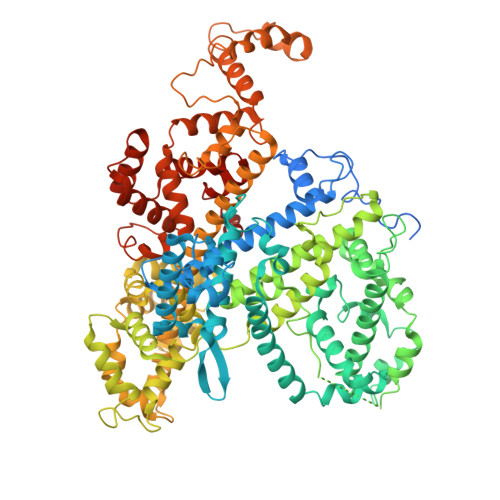
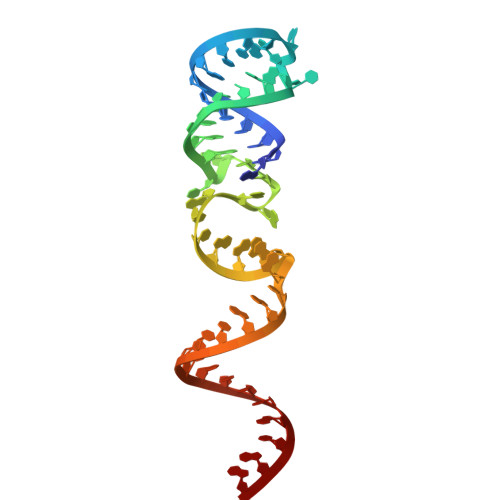
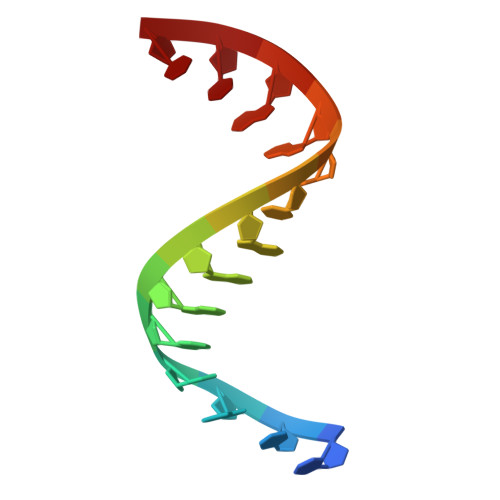
Cryo-EM structure of the RfxCas13d-crRNA-off-target-RNA complex.
Yang, Q., Sun, Y., Sun, L., Chi, T., Chen, Z.(2026) Structure 34: 11-19.e2
- PubMed: 41135510 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2025.09.010
- Primary Citation Related Structures:
9LZT, 9LZU - PubMed Abstract:
The CRISPR-Cas system is crucial for the adaptive immune response of prokaryotes and has been widely applied for genetic engineering. Cas13d, a type VI-D CRISPR-Cas effector, functions as RNA-guided ribonuclease and has been engineered for programmable RNA editing, which is a commonly used, active, and well-characterized small type VI editor. Here, we determined cryoelectron microscopy (cryo-EM) structures of Ruminococcus flavefaciens Cas13d in a RfxCas13d-crRNA-off-target-RNA ternary complex and RfxCas13d-crRNA binary complex at 3.10 and 3.13 Å resolution. The ternary complex consists of RfxCas13d, crRNA, and a captured short off-target ssRNA at a complex state of binding proximal mismatched RNA. RfxCas13d undergoes conformational changes with or without the off-target RNA, but the catalytic sites remain unchanged. Mg 2+ aids in stabilizing the crRNA repeat region structure, which may be crucial for RNA binding. This discovery provides the foundation for developing RfxCas13d as a mature tool and offers a framework for advancing transcriptome engineering.
- Shanghai Fifth People's Hospital and Institutes of Biomedical Sciences, Fudan University, Shanghai 200032, China.
Organizational Affiliation: