Cryo-EM structure of ligand-bound form of the receptor in complex with the transducer
Zhai, X., Guo, J., Shen, Q., Chen, L., Wang, G., Shen, D., Zhang, C., Xu, X., Mao, C., Zhang, Y., Liu, Z.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Beta-arrestin-1 | 393 | Bos taurus | Mutation(s): 0 Gene Names: ARRB1 | 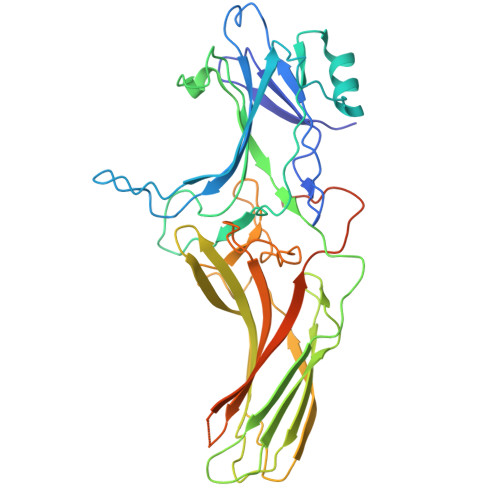 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P17870 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Fab30 heavy chain | B [auth H] | 229 | Mus musculus | Mutation(s): 0 | 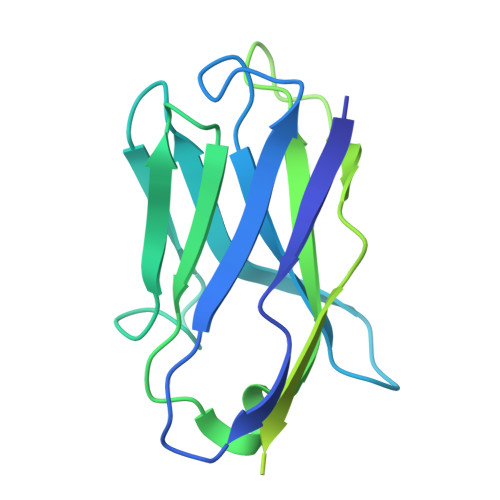 |
Entity ID: 3 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Fab30 light chain | C [auth L] | 215 | Mus musculus | Mutation(s): 0 | 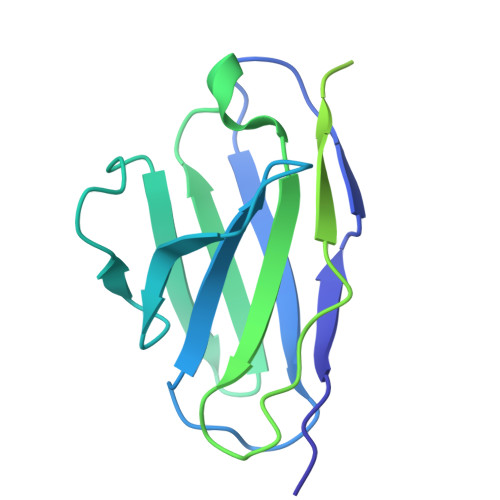 |
Entity ID: 4 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Long-acting PTH | D [auth P] | 36 | synthetic construct | Mutation(s): 0 | 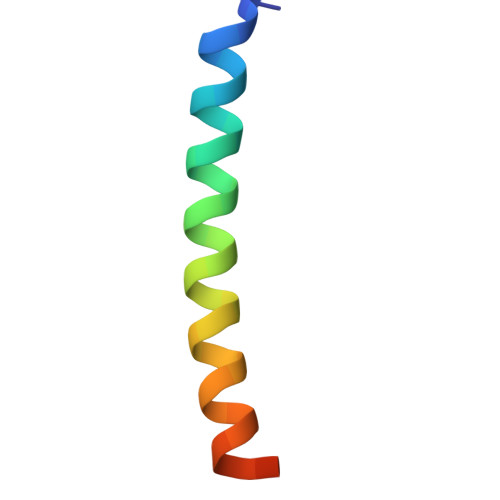 |
Entity ID: 5 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Parathyroid hormone/parathyroid hormone-related peptide receptor | E [auth R] | 483 | Homo sapiens | Mutation(s): 0 Gene Names: PTH1R, PTHR, PTHR1 | 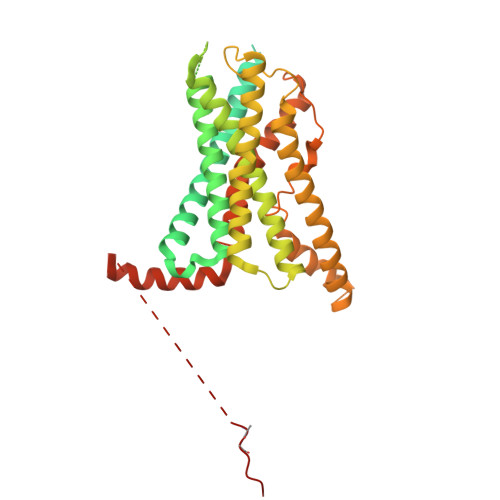 |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: Q03431 GTEx: ENSG00000160801 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q03431 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Modified Residues 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| SEP Query on SEP | E [auth R] | L-PEPTIDE LINKING | C3 H8 N O6 P |  | SER |
| TPO Query on TPO | E [auth R] | L-PEPTIDE LINKING | C4 H10 N O6 P |  | THR |
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | |
| RECONSTRUCTION | cryoSPARC |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Not funded | -- |