Structural basis for aerobic anoxygenic photosynthesis in the reaction center-light-harvesting 1 (RC-LH1) supercomplex of Dinoroseobacter shibae.
Liu, Z.K., Li, J.X., Zhang, Y.Y., Lv, J.L., Li, K., Chen, X.L., Zhang, Y.Z., Liu, L.N., Wang, P.(2025) Commun Biol 8: 1565-1565
- PubMed: 41238682 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-025-08935-7
- Primary Citation Related Structures:
9LTS, 9LTU, 9LTV - PubMed Abstract:
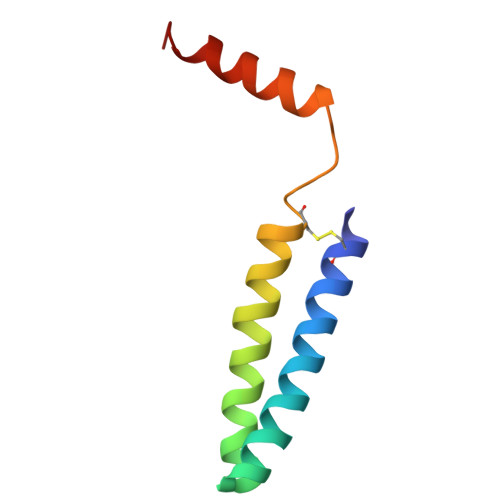
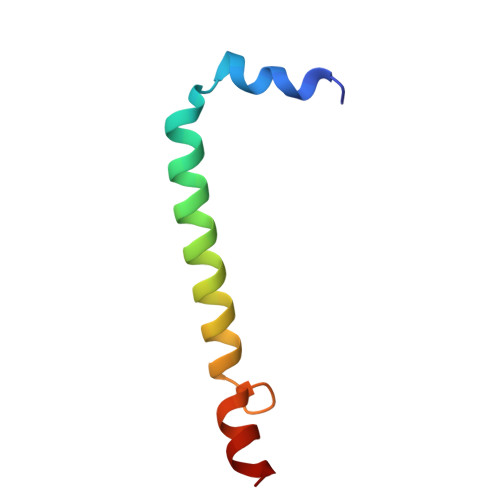
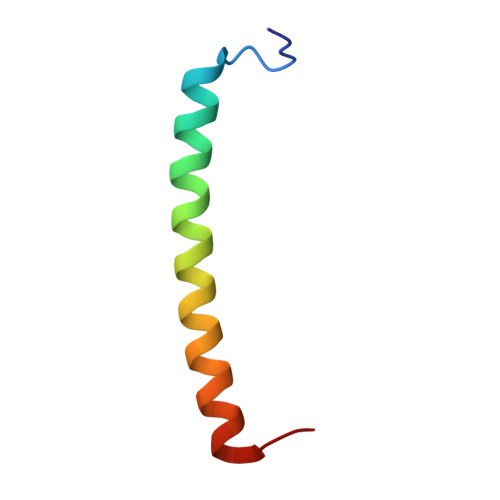
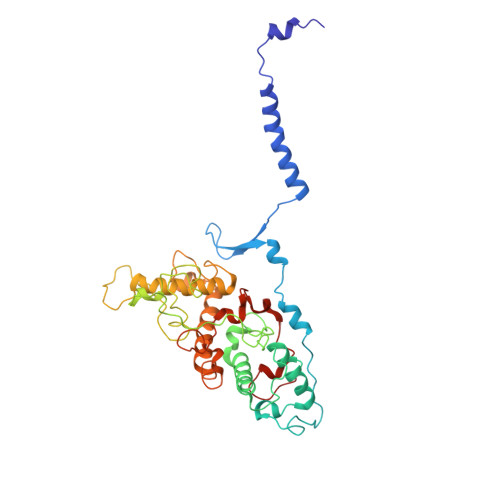
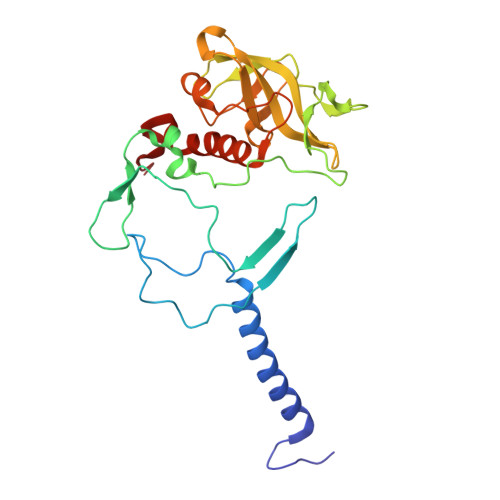
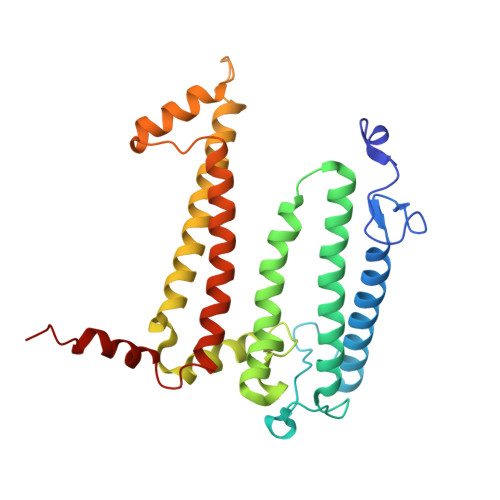
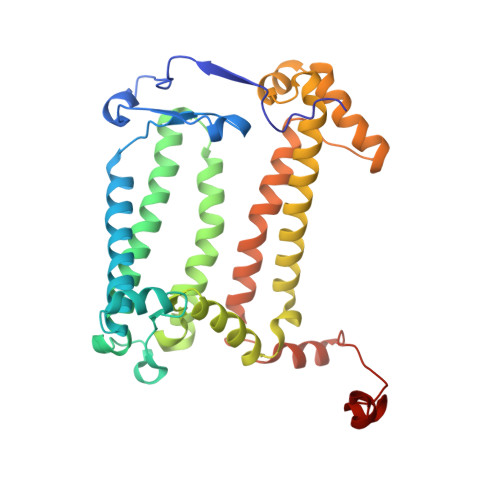
Aerobic anoxygenic phototrophic (AAP) bacteria are essential for oceanic carbon cycling. However, their architecture and structural adaptations of their photosynthetic systems to ensure adequate light harvesting, electron transport, and oxidative resilience in oxygen-rich environments remain poorly understood. In this study, we present a 2.4-Å cryo-EM structure of the reaction center-light-harvesting 1 (RC-LH1) supercomplex from Dinoroseobacter shibae DFL-12, a marine AAP bacterial symbiont of benthic dinoflagellates. This RC-LH1 supercomplex features a closed LH1 ring comprising 17 αβ-subunits, each containing two spheroidenones per αβ-heterodimer-a previously unreported configuration in phototrophic bacteria. The cytochrome subunit of the RC is truncated to three hemes, in contrast to the four-heme configuration found in anaerobic relatives. The structure also reveals elongated bacteriochlorophyll (BChl) spacing, which may account for its blue-shifted absorption maximum that is optimized for low-light benthic environments. Furthermore, we identify a previously unknown subunit, protein-LRC, which is hypothesized to functionally couple photochemical and respiratory electron transport. Collectively, these specific structural features allow AAP bacteria to balance anoxygenic photosynthesis and protection against oxidative damage, providing a mechanistic framework for them to thrive in oxygenated marine environments. Our study provides insights into the structural and functional variability of bacterial photosynthesis in response to oxygenated marine environments.
- MOE Key Laboratory of Evolution and Marine Biodiversity, State Key Laboratory of Marine Food Processing and Safety Control & College of Marine Life Sciences, Ocean University of China, Qingdao, China.
Organizational Affiliation: