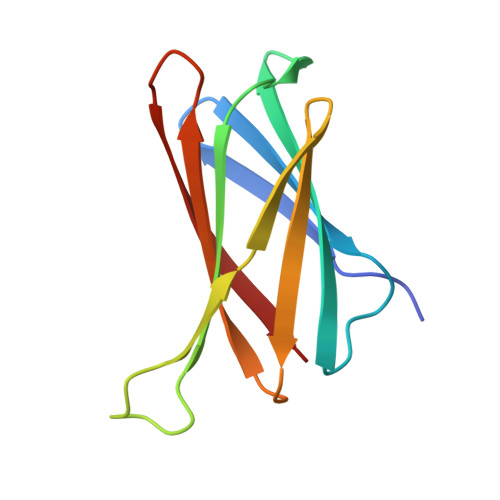
Structural basis of Cu(I) ion recognition by the Helicobacter pylori copper resistance determinant CrdA.
Ki, D.U., Oh, H.B., Cho, H.Y., Song, W.S., Yoon, S.I.(2026) FEBS J 293: 1785-1800
- PubMed: 41147750 Search on PubMed
- DOI: https://doi.org/10.1111/febs.70305
- Primary Citation Related Structures:
9LL2, 9LL4, 9LL5 - PubMed Abstract:
Helicobacter pylori is a bacterium that colonizes the stomach and causes gastric disorders in humans. For successful colonization in the harsh gastric environment, H. pylori employs various homeostatic mechanisms in response to environmental factors, such as protons and copper ions. Copper levels should be maintained below toxicity in the cell while remaining above the threshold required for biological functions. Copper resistance determinant A (CrdA) is a putative copper chaperone protein that contributes to copper homeostasis in H. pylori. To provide insight into CrdA-mediated copper homeostasis, we analyzed the interaction of CrdA with Cu(I) or Cu(II) ions through biochemical and mutational studies and determined the crystal structures of CrdA alone and in complex with a Cu(I) mimic, Ag(I). CrdA exhibited a binding preference for Cu(I) and Ag(I) ions over Cu(II) ions. CrdA forms a Greek key β-barrel structure with a unique protruding methionine-rich motif whose methionine residues coordinate two Ag(I) ions. In particular, the CrdA residue M69 plays a key role in Cu(I) recognition, even at the low pH found in the stomach where H. pylori resides. Furthermore, our structure-based comparative analysis suggests that CrdA has evolved a unique Cu(I) recognition mechanism that has not been observed for other Cu(I) chaperone proteins. Our findings on the specific interaction of CrdA with Cu(I) ions would provide a new avenue for developing H. pylori-targeting antibacterial drugs.
- Division of Biomedical Convergence, College of Biomedical Science, Kangwon National University, Chuncheon, Korea.
Organizational Affiliation: