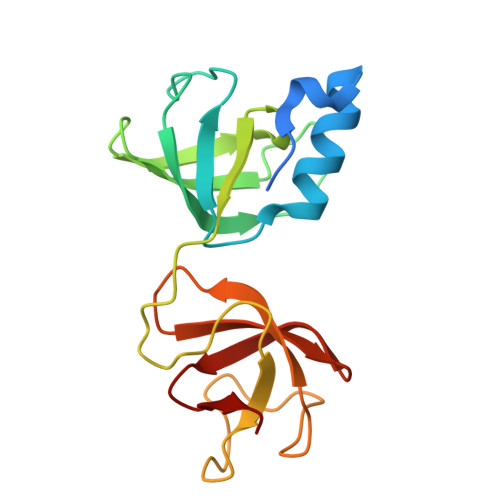
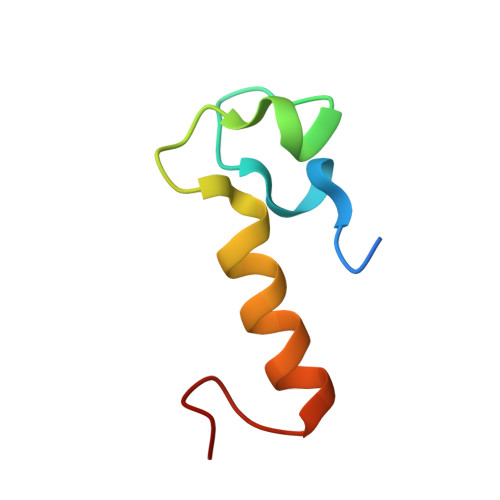
Mechanistic insights into the iron-sulfur cluster-dependent interaction of the autophagy receptor NCOA4 with the E3 ligase HERC2.
Liu, H., Shen, L., Gong, X., Zhou, X., Huang, Y., Zhou, Y., Guo, Z., Guo, H., Wang, S., Pan, L.(2025) Proc Natl Acad Sci U S A 122: e2510269122-e2510269122
- PubMed: 40705422 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.2510269122
- Primary Citation Related Structures:
9L8Y, 9L93 - PubMed Abstract:
NCOA4, a dedicated autophagy receptor for mediating selective autophagy of ferritin (ferritinophagy), plays a vital role in maintaining cellular iron homeostasis. The cellular abundance of NCOA4 is regulated by the E3 ligase HERC2 that can specifically target NCOA4 for proteasomal degradation under iron-replete conditions. However, the detailed molecular mechanism governing the iron-dependent recognition of NCOA4 by HERC2 remains elusive. Here, using multidisciplinary approaches, we systematically characterize the HERC2-binding domain (HBD) of NCOA4 and its interaction with HERC2. We uncover that NCOA4 HBD harbors a [2Fe-2S] cluster and can exist in two different states, the apo -form state and the [2Fe-2S] cluster-bound state. Moreover, we unravel that HERC2 can effectively recognize the [2Fe-2S] cluster-bound NCOA4 HBD through its Cullin-7-PARC-HERC2 (CPH) domain and iron-sulfur cluster-dependent NCOA4-binding domain (INBD) with a synergistic binding mode. The determined crystal structures of HERC2(2540-2700) and its complex with the [2Fe-2S] cluster-bound NCOA4 HBD together with relevant biochemical and cellular results not only elucidate how NCOA4 HBD specifically senses cellular iron level by binding a [2Fe-2S] cluster but also reveal the molecular basis underlying the specific interaction of HERC2 with the [2Fe-2S] cluster-bound NCOA4 HBD. In summary, our findings provide mechanistic insights into the iron-dependent turnover of NCOA4 by HERC2 and expand our understanding of the regulatory mechanism of NCOA4-mediated ferritinophagy.
- State Key Laboratory of Chemical Biology, Shanghai Institute of Organic Chemistry, University of Chinese Academy of Sciences, Chinese Academy of Sciences, Shanghai 200032, China.
Organizational Affiliation: