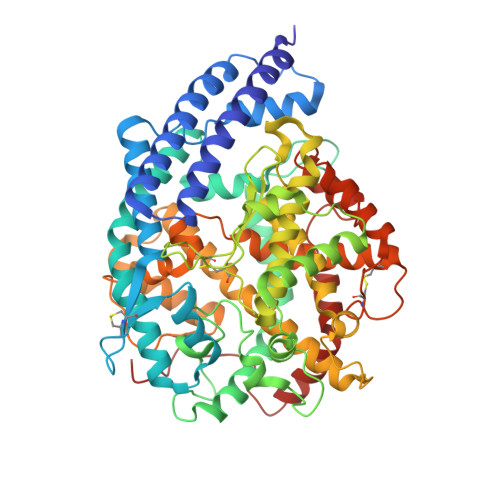
Cross-species recognition of squirrel ACE2 by the receptor binding domains of SARS-CoV-2, RaTG13, PCoV-GD and PCoV-GX.
Wang, C., Nan, X., Deng, Y., Fan, S., Lan, J.(2025) Structure 33: 1750
- PubMed: 40713966 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2025.07.003
- Primary Citation Related Structures:
9JR4, 9JR5, 9JR7, 9JRC, 9KUD - PubMed Abstract:
SARS-CoV-2 has a broad animal host range, but its origin and intermediate hosts are still debated. Arctic ground squirrel is proved to be a susceptible animal host of SARS-CoV-2 and RaTG13, as the Arctic ground squirrel ACE2 (sACE2) could bind to the RBDs of SARS-CoV-2 and RaTG13, and support transduction of the SARS-CoV-2 and RaTG13 pseudovirions. Here, we determined crystal structures of sACE2 bound to the RBDs of RaTG13, PCoV-GD, PCoV-GX, SARS-CoV-2 A372T and SARS-CoV-2 JN.1 variant. SPR assay indicated that RBD residues 493, 498 and 501 might be important for sACE2 recognition. Notably, N322-linked glycans of sACE2 were found to be in contact with S375 of the RaTG13 RBD. Moreover, the recent SARS-CoV-2 KP.2 and KP.3 variant RBDs could also bind to the sACE2. Our results provide further insights into interactions between coronavirus RBD and cell receptor ACE2, which may promote our understanding of SARS-CoV-2 evolution.
- School of Biomedical Sciences, Hunan University, Changsha, Hunan, China.
Organizational Affiliation: