Structural and Mechanistic Investigations of a High-Fidelity Cas9 from Parasutterella secunda
Shen, P., Lu, X., Wang, C., Huang, J.W., Min, J., Luo, J., Sun, Y., Chen, C.C., Guo, R.T.(2025) ACS Catal 15: 10109-10118
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
(2025) ACS Catal 15: 10109-10118
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| CRISPR-associated endonuclease Cas9 | 1,435 | Parasutterella secunda | Mutation(s): 0 | 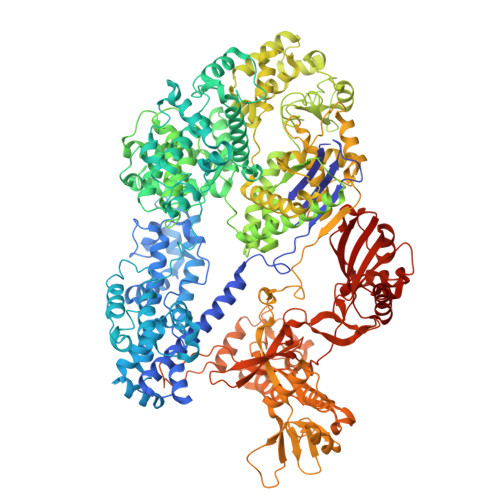 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | A0ABF7PQ28 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | ||||
| Molecule | Chains | Length | Organism | Image |
|---|---|---|---|---|
| RNA (131-MER) | 131 | Parasutterella secunda | 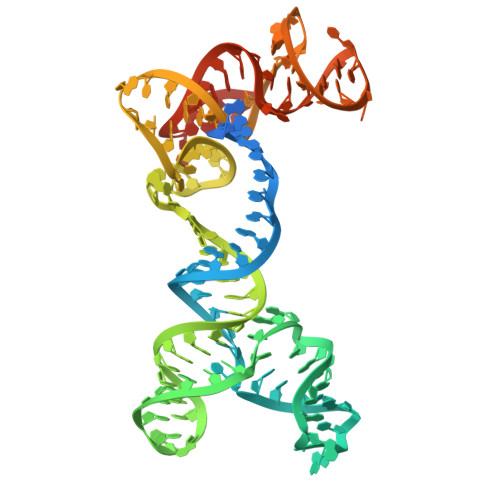 | |
Sequence AnnotationsExpand | ||||
Reference Sequence | ||||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Natural Science Foundation of China (NSFC) | China | -- |